Association of fecal and serum microRNA profiles with gastrointestinal cancer and chronic inflammatory enteropathy in dogs
Janne G. Lyngby, Marta Gòdia, and Louise Brogaard contributed equally as first authors.
Susanna Cirera and Lise N. Nielsen contributed equally as the last authors.
Funding information: Fondet for sygdomsbekæmpelse hos vore familiedyr (“Fund for Disease Control in our Companion Animals”), Grant/Award Number: A5386; The Independent Research Fund Denmark - Technology and Production, Grant/Award Number: DFF-6111-00124; The Danish Kennel Club
Abstract
Background
Reliable biomarkers to differentiate gastrointestinal cancer (GIC) from chronic inflammatory enteropathy (CIE) in dogs are needed. Fecal and serum microRNAs (miRNAs) have been proposed as diagnostic and prognostic markers of GI disease in humans and dogs.
Hypothesis/Objectives
Dogs with GIC have fecal and serum miRNA profiles that differ from those of dogs with CIE. Aims: (a) identify miRNAs that differentiate GIC from CIE, (b) use high-throughput reverse transcription quantitative real-time PCR (RT-qPCR) to establish fecal and serum miRNA panels to distinguish GIC from CIE in dogs.
Animals
Twenty-four dogs with GIC, 10 dogs with CIE, and 10 healthy dogs, all client-owned.
Methods
An international multicenter observational prospective case-control study. Small RNA sequencing was used to identify fecal and serum miRNAs, and RT-qPCR was used to establish fecal and serum miRNA panels with the potential to distinguish GIC from CIE.
Results
The best diagnostic performance for distinguishing GIC from CIE was fecal miR-451 (AUC: 0.955, sensitivity: 86.4%, specificity: 100%), miR-223 (AUC: 0.918, sensitivity: 90.9%, specificity: 80%), and miR-27a (AUC: 0.868, sensitivity: 81.8%, specificity: 90%) and serum miR-20b (AUC: 0.905, sensitivity: 90.5%, specificity: 90%), miR-148a-3p (AUC: 0.924, sensitivity: 85.7%, specificity: 90%), and miR-652 (AUC: 0.943, sensitivity: 90.5%, specificity: 90%). Slightly improved diagnostic performance was achieved when combining fecal miR-451 and miR-223 (AUC: 0.973, sensitivity: 95.5%, specificity: 90%).
Conclusions and Clinical Importance
When used as part of a diagnostic RT-qPCR panel, the abovementioned miRNAs have the potential to function as noninvasive biomarkers for the differentiation of GIC and CIE in dogs.
Abbreviations
-
- %CV
-
- coefficient of variance
-
- ANOVA
-
- analysis of variance
-
- AUC
-
- area under the curve
-
- CCECAI
-
- Canine Chronic Enteropathy Clinical Activity Index
-
- Cfa
-
- Canis familiaris
-
- CI
-
- confidence interval
-
- CIE
-
- chronic inflammatory enteropathy
-
- CPM
-
- counts per million
-
- CRP
-
- C-reactive protein
-
- Cq
-
- quantification cycle
-
- GI
-
- gastrointestinal
-
- GIC
-
- gastrointestinal cancer
-
- GIST
-
- gastrointestinal stromal tumor
-
- IBD
-
- inflammatory bowel disease
-
- LR
-
- likelihood ratio
-
- miRNA
-
- microRNA
-
- mRNA
-
- messenger RNA
-
- noPAP
-
- not containing poly(A) polymerase
-
- NPV
-
- negative predictive values
-
- NSAID
-
- nonsteroidal anti-inflammatory drug
-
- small RNAseq
-
- small RNA sequencing
-
- TLI
-
- trypsin-like immunoreactivity
-
- ROC
-
- receiver operating characteristic
-
- RT-qPCR
-
- high-throughput reverse transcription quantitative real-time PCR
-
- PLE
-
- protein losing enteropathy
-
- PPV
-
- positive predictive values
-
- UCPH
-
- University of Copenhagen
-
- WSAVA
-
- World Small Animal Veterinary Association
1 INTRODUCTION
Gastrointestinal cancer (GIC) is a heterogeneous group of diseases accounting for 8% of cancers in dogs.1 Clinical signs of GIC are nonspecific, including vomiting, diarrhea, weight loss, and anorexia, which makes it challenging to differentiate from other gastrointestinal (GI) disorders such as chronic inflammatory enteropathies (CIE). Though commonly part of the diagnostic work-up for enteropathy, abdominal ultrasonography has proven beneficial or vital in only 15% of dogs with chronic diarrhea, but having an abdominal or rectal mass increased the diagnostic utility.2 In some cases with solitary tumors or easy accessible lesions, minimally invasive cytological biopsies can be acquired. However, 1 study found that only 66% of cytological samples are clinically useful,3 and when clinically useful, cytology was in complete agreement with the histopathological diagnosis in 64% to 67% of the cases.3, 4 In diffuse neoplasia or even if a solitary tumor is visualized on ultrasound, endoscopic or surgical biopsies for histopathology might still remain necessary to achieve a diagnosis. This is costly, potentially invasive, and the required anesthesia carries an inherent risk. However, achieving a correct and timely diagnosis is in turn essential to instigate appropriate treatment, improve outcomes, and provide owners with a prognosis. Even with histology, some cancers such as small cell lymphoma can be challenging to distinguish from CIE, requiring additional diagnostics, delaying the time to diagnosis, and increasing the costs. Hence, there is a need for noninvasive biomarkers to reliably differentiate GIC from CIE in dogs before they undergo more invasive diagnostics.
A number of biomarkers for GIC have been investigated in human and veterinary medicine.5-10 MicroRNAs (miRNAs) have recently been proposed as promising diagnostic and prognostic markers of GI disease in both humans and dogs.11-16 MicroRNAs are small, non–protein-coding RNA molecules of approximately 20 to 23 nucleotide length, which primarily suppress the expression of target genes by binding to messenger RNA (mRNA) and inhibiting translation.17 MiRNAs have been detected in feces and serum from dogs and display substantial extracellular stability.14, 15 Aberrant serum miRNA profiles have been detected in numerous cancers in dogs,18 including intestinal T-cell lymphoma19 as well as in GI tissues of dogs with CIE.13, 19 However, the potential of miRNAs as noninvasive diagnostic fecal and serum biomarkers to differentiate GIC from CIE remains unknown.
We hypothesized that dogs with GIC display fecal and serum miRNA profiles that differ from those of healthy dogs and dogs with CIE, the aims were: (a) to identify relevant miRNAs by an initial global screening of the fecal and serum miRNAome in dogs using small RNA sequencing (small RNAseq) combined with a thorough investigation of human and veterinary literature to identify fecal and serum miRNAs with potential to differentiate between GIC, CIE, and healthy dogs. (b) To use high-throughput reverse transcription quantitative real-time PCR (RT-qPCR) to establish a fecal and serum miRNA panel with the potential to distinguish GIC from CIE in dogs.
2 MATERIALS AND METHODS
2.1 Study design and ethics approval
This international multicenter observational prospective case-control study was carried out from 2018 to 2020 and initiated by the University of Copenhagen (UCPH). Participating centers included: Norwegian University of Life Sciences, University of Glasgow, University of Edinburgh. Each institution obtained their own local ethical approval (#2017-9 + 2017-12; 14/04723-72; Ref 05a/18; VERC#41.18). In addition, the Animal Experiments Inspectorate under the Ministry of Food, Agriculture, and Fisheries of Denmark, Danish Veterinary and Food Administration (case #2017-15-0201-01353) approved the study. Owners provided informed written consent for study participation and could withdraw this at any time.
Dogs were recruited at UCPH, the Norwegian University of Life Sciences, University of Glasgow, the University of Edinburgh, or at Evidensia Karlslunde Animal Hospital, or Evidensia Faxe Animal Hospital. All healthy dogs were recruited at UCPH.
2.2 Dog recruitment
Client-owned dogs were prospectively recruited for the study and allocated into 1 of 3 groups: GIC, CIE, and healthy dogs (Table S1). Dogs presenting for evaluation of chronic (>2-3 weeks duration) signs of GI disease, suspicion of GIC, or both, and undergoing endoscopic or surgical biopsies were enrolled.
Dogs in all groups had to be ≥1 year of age and have a bodyweight ≥2.5 kg. Dogs were not included if they were being fed a raw food diet, had received antimicrobial therapy, corticosteroids, or other immunomodulatory drugs within the last 6 weeks, or nonsteroidal anti-inflammatory drugs within the 2 weeks before enrolment. All 3 groups of dogs underwent a standardized minimum diagnostic work-up including a history, physical exam, complete blood count, serum biochemistry, serum C-reactive protein (CRP) concentration, serum folate, and cobalamin concentrations, urinalysis, fecal flotation (Fecalyzer, Vetoquinol, Lure, France), and Giardia (Immunocard STAT! Crypto/Giardia, Meridian Bioscience, San Diego, California, USA) testing. The Canine Chronic Enteropathy Clinical Activity Index (CCECAI)20 was calculated and a fecal score was determined using the Purina 7-point fecal scoring system (Nestlé S.A., Vevey, Switzerland).21, 22
In dogs suspected of CIE, additional diagnostic procedures were performed including but not limited to serum basal cortisol concentration, ACTH stimulation test if basal cortisol ≤55 nmol/L,23 and serum trypsin-like immunoreactivity (TLI) concentration to rule out other GI and non-GI diseases.24, 25 All CIE dogs underwent a dietary trial with a hydrolyzed or novel protein diet for a minimum of 14 days, and were excluded if they improved.12 Dogs with antibiotic-responsive enteropathy were likewise excluded; however, because of a focus on prudent antibiotic use and strict Danish antimicrobial guidelines for dogs recruited in Denmark, an antibiotic trial was not routinely undertaken.
Further exclusion criteria for the initial small RNAseq screening were the presence of concurrent diseases, serum albumin <2.0 g/dL (<20 g/L), or histopathological evidence of lymphangiectasia. For RT-qPCR analysis, comorbidities were permissible in dogs with GIC, and treatment with antibiotics, glucocorticoids, or other medication was allowed. Abdominal imaging (ultrasound or computed tomography scan) was performed in all GIC or CIE dogs and thoracic imaging was encouraged. Histopathology including World Small Animal Veterinary Association (WSAVA) scoring of intestinal biopsies was available for all cases of CIE.26, 27
Healthy client-owned dogs were enrolled based on history and physical exam and no episodes of signs of GI disease within the last 2 months. As part of a meticulous screening to ensure a clear phenotypical characterization the healthy dogs enrolled in the small RNAseq screening underwent an upper and lower GI endoscopy under general anesthesia with mucosal pinch biopsies from the stomach, duodenum, ileum, and colon.
2.3 Sample collection
Blood samples were collected at presentation and processed within 30 minutes of sampling. Serum for miRNA analysis was aliquoted and stored at −20°C. Fecal samples were collected and aliquoted (approximately 1 g per aliquot) into cryotubes within 1 hour of defecation and stored at −20°C for miRNA quantification. This was done at the presentation or at home by the client after receiving thorough verbal and written instructions and the necessary supplies as previously described.15, 28 Mucosal endoscopic pinch biopsies or surgical full-thickness biopsies were obtained at the clinician's discretion. Histopathologic tissue evaluation was performed by a reference laboratory, and inflammatory changes were evaluated using the WSAVA Standardization Grading System.26, 27 All frozen samples were stored at −80°C for batch analysis at a later stage.
2.4 RNA isolation
For fecal samples, total RNA was isolated from 100 mg fecal material using the miRNeasy Mini Kit (Qiagen, Hilden, Germany) as described previously.28
For serum samples, 200 μL were used for isolation of total RNA using the miRNeasy Serum/Plasma kit (Qiagen, Hilden, Germany) without including the accompanying Caenorhabditis elegans miR-39 spike-in extraction control.
Concentration and purity of each RNA sample were determined by assessing the 260/280 and 260/230 absorbance ratios on a NanoDrop ND-1000 Spectrophotometer (Thermo Scientific, Hvidovre, Denmark), and stored at −80°C until further use.
2.5 Small RNAseq and data analysis
Small RNAseq libraries were prepared using the NEBNext Small RNA Library Prep Set for Illumina (New England Biolabs, Ipswich, Massachusetts) using 6 μL total RNA from each fecal or serum sample. This was carried out by the sequencing service provider (Genomics Unit, Center for Genomic Regulation, Barcelona Biomedical Research Park, Barcelona, Spain) using the Illumina HiSeq 2500 system to produce 50 bp single-end reads. A fully detailed description of small RNAseq data analysis can be found in File S1.29-37
2.6 Validation of candidate miRNAs using RT-qPCR
Identification of GIC biomarker candidates was performed using RT-qPCR. The high-throughput qPCR platform Biomark HD (Fluidigm, San Francisco, California) was applied for this purpose in combination with the appertaining 96.96 Dynamic Array integrated fluidic circuit chips (Fluidigm, San Francisco, California). A fully detailed description of RT-qPCR can be found in File S1.
2.7 Data analyses
2.7.1 Clinical data
Data with continuous variables were assessed for normality using the Shapiro-Wilk test. Summary statistics for continuous and ordinal variables were reported as median and range and categorical data were presented as absolute and relative frequencies (number, n, and percentage, %). The CCECAI score was considered an ordinal variable. Continuous variables were compared using an unpaired Student's t test or Mann-Whitney test when comparing 2 groups, and a 1-way analysis of variance (ANOVA) or Kruskal-Wallis test when comparing 3 groups. Categorical data were compared using Fisher's exact test or Chi-squared test. Bonferroni's multiple comparisons test or false discovery rate using the Benjamini, Krieger, and Yekutieli method were used to correct for multiple comparisons. For all statistics, a P value < .05 was considered significant. Statistical software packages (GraphPad Prism 9.2.0) were used for all statistical analyses and preparation of graphs.
2.7.2 RT-qPCR data
Inspection of amplification curves, melting curves, and standard curves as well as negative controls (noPAP and nontemplate controls) was performed using the Fluidigm Real-Time PCR Analysis software (v. 4.7.1, Fluidigm). GenEx Pro software (v. 7.1.1.118, MultiD Analyses AB) was used for interplate calibration, qPCR efficiency correction of Cq values, evaluation of endogenous miRNA normalizers using the algorithms NormFinder38 and geNorm,39 normalization with the most suitable normalizers, calculation of relative quantities, and statistical analysis of miRNA amounts between groups. When comparing 3 groups, statistical significance of the difference in miRNA amounts was assessed with 1-way ANOVA and Tukey-Kramer's test after pairwise comparisons. When comparing miRNA amounts in only 2 groups, Student's t test was applied. Significance levels were set to ɑ = .0016 (feces) and ɑ = .00070 (serum) to account for performing multiple comparisons (Šidàk corrected P values, as employed by the GenEx Pro software). Analyses yielding P values between .05 and the corrected P values were interpreted with caution. MiRNA fold changes were computed as the ratio between the average relative miRNA amounts in the 2 groups being compared, and with a 95% confidence interval.
Correlation of miRNA quantification as determined by small RNAseq and RT-qPCR was assessed using Pearson's correlation coefficient (Pearson's r) using GraphPad Prism.
The ability of selected miRNAs to discriminate between GIC and CIE dogs was assessed by receiver operating characteristic (ROC) curve analysis using GraphPad Prism. Diagnostic performance of individual miRNAs or combinations of miRNAs was evaluated by determining sensitivity and specificity as well as positive and negative likelihood ratios (LR+, LR−). To this end, the optimal cutoff level for each miRNA was calculated using the maximum value of Youden's index. Multiple logistic regression was applied to investigate the combined effect of multiple miRNAs for discrimination between GIC and CIE dogs using GraphPad Prism.
3 RESULTS
3.1 Dog characteristics
The GIC group consisted of 24 dogs (median age: 101 months, [range, 27-192 months]; 9 females/15 males). The most common breeds were Eurasians (n = 4), French bulldogs (n = 2), and mix breed dogs (n = 3). A complete list of breeds can be found in Table S1. Eleven dogs had hematemesis, melena or hematochezia. Fourteen dogs had an endoscopy, 7 dogs had a laparotomy, and additionally, 3 dogs initially had an endoscopy but because of concern of not having the correct diagnosis, they underwent a laparotomy with full thickness biopsies; these 3 dogs all had small or intermediate cell lymphoma. Final diagnoses included: lymphoma (n = 9/24; small cell lymphoma n = 3, intermediate cell lymphoma n = 1, large cell lymphoma n = 5; gastric n = 2, small intestinal n = 7), carcinoma (n = 10/24; gastric n = 8, small intestinal n = 1, rectal n = 1), and sarcomas (n = 4/24; gastrointestinal stromal tumor [GIST] n = 3, nonclassified n = 1). Sarcomas were located in the stomach (n = 1) and small intestine (n = 3). One dog was diagnosed with a GI mast cell tumor (heavy tumor burden in the stomach and duodenum). Twelve dogs had comorbidities, including cardiac disease or hypothyroidism, or had received glucocorticoids, antibiotics, NSAIDs, or a combination thereof, before enrolment. However, when comparing dogs with comorbidities or therapeutic intervention to dogs without, there was no statistical significance in age (P = .09), body weight (P = .11), sex (P = .67), duration of disease (P = .64), clinical signs (P ≥ .99), CCECAI score (P = .64), serum cobalamin concentration (P = .7), serum folate concentration (P = .76), or serum CRP concentration (P = .87). These 24 dogs are described as 1 GIC group throughout the article.
Ten dogs with CIE were enrolled. All dogs had clinical signs of small intestinal disease alone or together with gastritis or colitis. Dogs with CIE had a longer duration of disease (median: 284 days, range, 45-730 days) compared to GIC (median: 120 days, range, 3-730 days; P = .02). The CCECAI score was significantly different with a median of 6 (range, 3-12) in CIE and 11 (1-17) in GIC (P = .04), but the distribution of clinical signs did not differ between CIE and GIC groups (Table 1). All dogs had serum resting cortisol concentration measured, 5/10 had an ACTH stimulation test, 9/10 had serum TLI concentration measured, and 5/10 had serum pancreatic lipase immunoreactivity concentration measured. Four dogs had hematemesis, melena, hematochezia, or a combination hereof. All dogs had an upper and lower endoscopy performed and the ileum was accessed in 8 of the 10 dogs. The dominant histopathological diagnoses were lymphocytic-plasmacytic enteritis (n = 7), gastritis and colitis (n = 1), gastritis and enteritis (n = 1), or gastritis, enteritis, and colitis (n = 1). The 1 dog with gastritis combined with colitis as the dominant histopathological diagnosis did also have mild enteritis as well as clinical signs of small intestinal disease.
| Group characteristic | Healthy dogs | CIE | GIC | P valuea |
|---|---|---|---|---|
| Total number, n | 10 | 10 | 24 | |
| Dog characteristics | ||||
| Age in monthsb, median (range) | 61 (15-130) | 53 (18-132) | 101 (27-192) | .02 |
| Sex, n, female/male | 5/5 | 3/7 | 9/15 | .87 |
| Body weight in kgc, median (range) | 14 (6.5-33) | 10 (2.6-22) | 15 (2.4-74) | .31 |
| Clinical variables | ||||
| Disease duration in daysc, median (range) | NA | 284 (45-730) | 120 (3-730) | .02 |
| Clinical signs, n (%) | ||||
| Vomiting | NA | 5/10 (50%) | 19/23 (83%) | .09 |
| Diarrhea | NA | 7/10 (70%) | 14/23 (61%) | .71 |
| Weight loss | NA | 3/9 (33%) | 14/23 (61%) | .24 |
| Hyporexia/ anorexia | NA | 8/10 (80%) | 21/23 (91%) | .57 |
| GI bleeding (hematemesis, hematochezia, or melena) | NA | 4/10 (40%) | 11/24 (46%) | >.99 |
| CCECAI scored, median (range) | 1 (0-4) | 6 (3-12) | 11 (1-17) | .04 |
| Clinical disease severity, n (%) | 10/10 (100%) | 9/10 (90%) | 18/24 (75%) | .33 |
| Mild (CCECAI score ≤5) | 10 (100%) | 3 (33%) | 2 (11%) | |
| Moderate (CCECAI score 6-8) | 0 | 3 (33%) | 4 (22%) | |
| Severe (CCECAI score 9-11) | 0 | 1 (11%) | 4 (22%) | |
| Very severe (CCECAI score ≥ 12) | 0 | 2 (22%) | 8 (45%) | |
| Clinicopathological variables | ||||
| Serum cobalamine (pmol/L), median (range) | 341 (242-1066) | 311 (145-573) | 204 (105-675) | .06 |
| Serum folatef (nmol/L), median (range) | 27.9 (20.5-54.4) | 24.3 (6.2-93.1) | 16.7 (2.5-50.8) | .18 |
| CRPf (g/L), median (range) | 5.7 (0-7.4) | 6.8 (0.7-10.2) | 14.1 (0-104.7) | .15 |
- Note: All 3 study groups are compared in the statistical analyses except of “Clinical variables” as this was not relevant for the healthy dogs.
- Abbreviations: CCECAI, canine chronic enteropathy clinical activity index; CIE, chronic inflammatory enteropathy; CRP, C-reactive protein; GIC, gastrointestinal cancer; GI, gastrointestinal.
- a Significant difference between groups (P < .05) is indicated with boldface values.
- b Significant difference was observed when comparing all 3 groups in an ANOVA; the difference remains significant only between GIC and CIE when using Bonferroni's multiple comparisons test (P = .04).
- c One dog was censored because of missing data.
- d Significant difference between GIC and CIE dogs using Mann–Whitney test. Statistical comparisons to the healthy group were not made for the CCECAI score. Seven dogs were censored because of missing values in CCECAI.
- e No significant difference was observed when comparing all 3 groups in a Kruskal–Wallis test; however, there was a significant difference between GIC and healthy dogs when the false discovery rate using the Benjamini, Krieger, and Yekutieli method was applied post hoc (P = .02). Not documented in 6 dogs.
- f Not documented in 5 dogs.
3.1.1 RNA isolation
Of the 44 enrolled dogs, feces were obtained from 42 dogs and serum was obtained from 41 dogs, to be used for RNA isolation (Figure 1, Table S1). Yield of RNA from fecal samples ranged from 27.9 to 864.1 ng/μL (average 282.6 ng/μL) with average 260/280 and 260/230 ratios of 1.98 and 1.37, respectively. Yield of RNA from serum samples ranged from 7.5 to 49.6 ng/μL (average 16.5 ng/μL) with mean 260/280 and 260/230 ratios of 1.45 and 0.38, respectively.
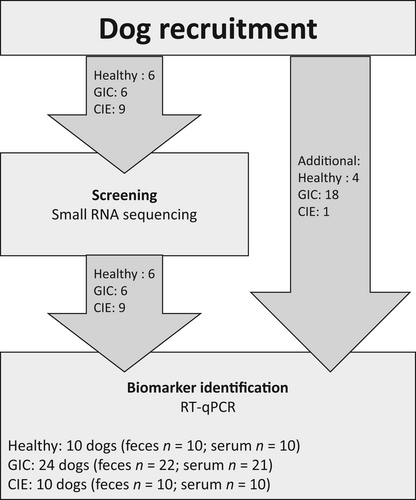
3.1.2 Small RNAseq screening of feces and serum from dogs
A total of 21 dogs (6 healthy dogs, 9 dogs with CIE, and 6 dogs with GIC) were included in the initial small RNAseq screening (Figure 1, Table S1). Small RNAseq libraries yielded an average of 11.2 M and 10.9 M reads for fecal and serum samples, respectively. Over 98% of reads passed quality control filters (Table S3). On average, 24% and 84% of the filtered reads mapped to the Canis familiaris (cfa) genome in fecal and serum samples, respectively (Table S3). Of these mapped reads, 0.02% in feces and 4.3% in serum mapped to annotated mature miRNAs in the C familiaris genome (Table S3; Figure S1). Of the 453 mature C familiaris miRNAs annotated in miRBase (v.22.1) a total of 210 and 283 miRNAs were identified in at least 1 fecal or serum sample, respectively (Table S4). On average, the most abundant miRNAs in feces were cfa-miR-21 (177 930 counts per million [CPM]), cfa-miR-192 (104 096 CPM), and cfa-miR-215 (99 857 CPM), whereas the most abundant miRNAs in serum were cfa-miR-486-3p (286 956 CPM), cfa-miR-486 (263 105 CPM), and cfa-miR-92a (63 269 CPM; Table S4).
By setting a threshold of a q value <.05 and a |log2FC| ≥1.5, 1 fecal miRNA (cfa-miR-451) and 5 serum miRNAs (cfa-miR-133c, cfa-miR-133a, cfa-miR-145, cfa-miR-143, cfa-miR-1) were found to be present at significantly different amounts in GIC and CIE dogs, and all 6 miRNAs were present at higher amounts in GIC compared to CIE (Figure S2, Table S5). Comparison of GIC and healthy dogs revealed 2 additional fecal miRNAs (cfa-miR-194, lower in GIC, and cfa-miR-320, higher in GIC) and 1 serum miRNA (cfa-miR-122, higher in GIC) to be different (Figure S2, Table S5). When comparing CIE and healthy dogs, no differences were found.
3.1.3 RT-qPCR identification of GIC biomarkers
Of the 96 miRNAs included in the panel, 36, and 75 assays were successfully quantified using RT-qPCR in feces and serum, respectively (Tables S240-57 and S6). Successful quantification was determined based on the absence of or negligible fluorescence in noPAP controls, mono-peaked melting curves indicating specific amplification of the desired product, acceptable qPCR efficiency (ranged from 93% to 109% for feces, 91% to 114% for serum, Table S2),40-57 and good cDNA reproducibility (less than 1 Cq difference between cDNA replicates). miR-16, miR-29a, and miR-148b were used for the normalization of fecal data, and miR-148b and miR-320 were used for the normalization of serum data.
Significantly higher relative quantity of miR-451 were detected in the feces of dogs with GIC compared to CIE, confirming results obtained in the small RNAseq screening (Table 2, Figure 2A). Additionally, RT-qPCR results demonstrated significantly higher miR-27a and miR-223 amounts in the feces of dogs with GIC compared to CIE (Table 2, Figure 2A). Increased amounts of miR-143-3p and miR-145 in serum of GIC dogs compared to CIE as demonstrated in the small RNAseq screening were confirmed by RT-qPCR, although there was no statistical significance after correcting for multiple testing (Table 2, Figure 2B). RT-qPCR also demonstrated significantly increased serum amounts of miR-20b, miR-148a-3p, and miR-652 in GIC compared to CIE (Table 2, Figure 2B).
| P value (1-way ANOVA) | FC (GIC vs CIE) (±95% CI) | FC (GIC vs healthy) (±95% CI) | FC (CIE vs healthy) (±95% CI) | Significant correlation between small RNAseq and qPCR? (Pearson's r) | |
|---|---|---|---|---|---|
| Fecal miRNAs | |||||
| miR-451 | <.001 | 59 (±38) | 59 (±38) | 0.99 (±0.63) | Yes, P < .001, r = .78 |
| miR-223 | <.001 | 25 (±13) | 19 (±10) | 0.76 (±0.58) | Nob, P = .09, r = .53 |
| miR-27a | <.001 | 2.3 (±0.40) | 2.1 (±0.37) | 0.93 (±0.31) | Nob, P = .51, r = .18 |
| Serum miRNAs | |||||
| miR-20b | <.001 | 2.3 (±0.53) | 2.6 (±0.61) | 1.2 (±0.21) | Yes, P = .003, r = .62 |
| miR-148a-3p | <.001 | 2.2 (±0.39) | 2.2 (±0.39) | 1.0 (±0.16) | Yes, P < .001, r = .75 |
| miR-652 | <.001 | 5.0 (±2.2) | 3.7 (±1.7) | 0.74 (±0.21) | Yes, P < .001, r = .77 |
| miR-145 | .0034a | 5.0 (±3.1) | 4.9 (±3.1) | 0.98 (±0.23) | Yes, P < .001, r = .89 |
| miR-143-3p | .0075a | 5.3 (±3.1) | 3.1 (±1.8) | 0.59 (±0.12) | Yes, P < .001, r = .86 |
- Abbreviations: ANOVA, analysis of variance; CIE, chronic inflammatory enteropathy; CI, confidence interval; FC, fold change; GIC, gastrointestinal cancer.
- a P value below .05, but does not survive correction for multiple comparisons in 1-way ANOVA.
- b Not quantifiable in several samples with small RNAseq (48% missing values for miR-223; 28% missing values for miR-27a). Fold change value in bold: P value for this pairwise comparison (Tukey–Kramer's test) survives correction for multiple comparisons. Fold change value underlined: P value below .05 for this pairwise comparison (Tukey–Kramer's test), but does not survive correction for multiple comparisons. All other comparisons: nonsignificant. 95% CI for fold changes are indicated in brackets.
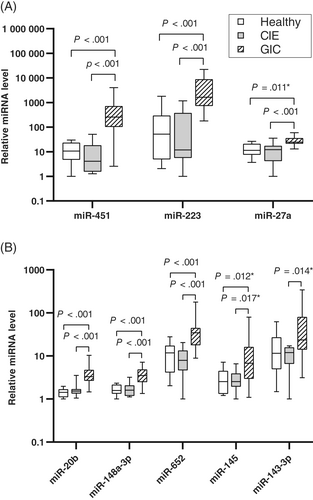
No significant differences in any fecal or serum miRNA amounts were demonstrated when comparing CIE to healthy dogs (Table 2, Table S6). Within the GIC group, having comorbidities or having received glucocorticoids, antibiotics, or NSAIDs did not affect miRNA amounts (Table S6). Also, across different types or locations of GIC, no significant differences in miRNA amounts were observed (Table S6).
3.1.4 Diagnostic performance of fecal and serum miRNAs
When assessing miRNAs for their ability to discriminate between GIC and CIE, the best performances were seen for fecal miR-451, miR-223, and miR-27a, and serum miR-20b, miR-148a-3p, and miR-652, with an area under the curve (AUC) of 0.955, 0.918, and 0.868, and 0.905, 0.924, and 0.943, respectively (Figure 3, Table 3). Combining miRNAs resulted in slight improvements in diagnostic performance, with the greatest improvement observed for the combination of fecal miR-451 and miR-223. This combination of markers yielded an AUC of 0.973, a sensitivity of 95.5%, a specificity of 90%, and LR+ and LR− of 9.55 and 0.05, respectively (Table 3).
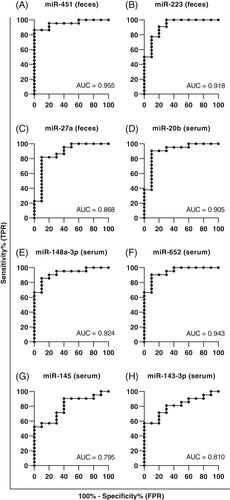
| AUC | 95% CI (AUC) | Sn (%) | 95% CI (Sn) | Sp (%) | 95% CI (Sp) | LR+ | LR− | |
|---|---|---|---|---|---|---|---|---|
| Fecal miRNAs | ||||||||
| miR-451 | 0.955 | 0.888-1.0 | 86.4 | 66.7-95.3 | 100 | 72.3-100 | Infinity | 0.14 |
| miR-223 | 0.918 | 0.810-1.0 | 90.9 | 72.2-98.4 | 80 | 49.0-96.5 | 4.55 | 0.11 |
| miR-27a | 0.868 | 0.715-1.0 | 81.8 | 61.5-92.7 | 90 | 59.6-99.5 | 8.18 | 0.20 |
| miR-451 + miR-223 | 0.973 | 0.925-1.0 | 95.5 | 78.2-99.8 | 90 | 59.6-99.5 | 9.55 | 0.05 |
| Serum miRNAs | ||||||||
| miR-20b | 0.905 | 0.778-1.0 | 90.5 | 71.1-98.3 | 90 | 59.6-99.5 | 9.05 | 0.11 |
| miR-148a-3p | 0.924 | 0.830-1.0 | 85.7 | 65.4-95.0 | 90 | 59.6-99.5 | 8.57 | 0.16 |
| miR-652 | 0.943 | 0.863-1.0 | 90.5 | 71.1-98.3 | 90 | 59.6-99.5 | 9.05 | 0.11 |
| miR-145 | 0.795 | 0.636-0.955 | 52.4 | 32.4-71.7 | 100 | 72.3-100 | Infinity | 0.48 |
| miR-143-3p | 0.810 | 0.659-0.960 | 57.1 | 36.6-75.5 | 100 | 72.3-100 | Infinity | 0.43 |
- Note: Diagnostic performance of miRNAs with regards to distinguishing between GIC and CIE. For any analyses including only fecal miRNA: n = 32 (22 GIC, 10 CIE); for any analyses including only serum miRNA: n = 31 (21 GIC, 10 CIE).
- Abbreviations: AUC, area under the curve; CI, confidence interval; LR−, negative likelihood ratio; LR+, positive likelihood ratio; Sn, sensitivity; Sp, specificity.
4 DISCUSSION
This study demonstrates an association between the miRNAome in feces and serum in clinically well-characterized dogs with either GIC or CIE. Fecal miR-451, miR-223, and miR-27a, as well as serum miR-20b, miR-148a-3p, and miR-652, were identified as noninvasive potential biomarkers for the differentiation of GIC and CIE by quantifying a comprehensive panel of miRNAs in feces and serum using RT-qPCR. Upon validation in a larger and clinically more diverse group of dogs with chronic GI disease, future application of some or all of these miRNAs in a diagnostic qPCR assay might facilitate early and timely differentiation between GIC and CIE in dogs before more invasive diagnostics are performed.
Results demonstrate miR-451 and miR-223 to be robust fecal biomarkers for GIC in dogs, which mirrors findings regarding colorectal cancer in people.58, 59 miR-451 is highly expressed in erythrocytes and miR-223 is highly abundant in, for example, neutrophils.60, 61 The increase in miR-451 and miR-223 amounts observed in feces of GIC dogs could therefore reflect increased presence of blood in the feces of dogs with GIC compared to CIE. For this same reason, these 2 miRNAs are suggested as specific fecal biomarkers of necrotizing enterocolitis in preterm infants.62 Consequently, whether the presence of miR-451 and miR-223 in feces from dogs is unambiguously indicative of GIC should be addressed in future studies, as other GI diseases might likewise cause blood to be present in the feces. Investigating fecal miRNA amounts in relation to intestinal bleeding was not an aim of the present study; however, we subsequently identified the occurrence of intestinal bleeding (melena, hematochezia, hematemesis, or a combination thereof) to be comparable in the GIC and CIE groups. Neutrophil influx during intestinal inflammation should also be considered as a source of miR-223, in case these cells are shed or leaked into feces even in the absence of intestinal bleeding.63
miR-27a is generally described as an oncomiR (miRNA oncogene).64 High amounts of miR-27a in human colorectal cancer have been shown to support cell proliferation and tumor growth and to be associated with resistance toward chemotherapy.65 The promotion of cancer cell proliferation and metastasis has likewise been described in relation to gastric cancer.66, 67 In contrast to miR-451 and miR-223, the observed increase of miR-27a in feces from dogs with GIC is more likely to be a direct effect of up-regulation in cancerous tissue, as occurs in humans with colorectal cancer11, 68 and gastric cancer,68, 69 and might therefore be a more reliable GIC biomarker if no GI bleeding is present, as the increase of fecal miR-27a amounts in GIC dogs would not rely on GI bleeding. However, the role of miR-27a in GIC is not currently clear, and further investigations are warranted in order to confirm its oncomiR function in dogs. The varying fold change amounts that we observe between GIC and CIE dogs also support the different mechanisms suggested for the observed increased fecal amounts of miR-451, miR-223, and miR-27a in GIC dogs. Here, we observed a 2.3-fold increase in fecal miR-27a amounts in GIC dogs compared to CIE, whereas the increase for miR-451 and miR-223 was 59- and 25-fold, respectively. This discrepancy suggests a difference in the underlying mechanisms resulting in the presence of these miRNAs in the feces of GIC dogs, supporting that the slight increase in miR-27a amounts could be because of its up-regulation locally in tumor tissue and after release into feces, whereas the substantial increase of miR-451 and miR-223 is likely because of increased amount of blood in feces.
Should fecal miR-27a amounts indeed be a reflection of local tumor up-regulation of expression, then it could potentially not be equally well suited as a biomarker for all types of GIC in dogs. Although we did not observe any significant differences in miR-27a amounts when cancer dogs were grouped according to cancer type (carcinoma, lymphoma, sarcoma), we did however find that miR-27a was inferior to miR-451 and miR-223 in discriminating between GIC and CIE dogs; in this regard, the likely blood-derived biomarkers miR-451 and miR-223 performed more robustly.
Fecal samples proved challenging to collect in several dogs, because dogs with prolonged anorexia or hyporexia can have decreased fecal volume. In such dogs, serum biomarkers would prove more suitable, but in the majority of dogs, feces is easily accessible and the ideal noninvasive matrix for GI disease. From a technical perspective, obtaining successful quantification of miRNAs with RT-qPCR proved challenging using feces. The same 96 assays were applied to both fecal and serum samples, and whereas 75 assays were successful in serum, only 36 achieved acceptable results in feces. The primary issue appeared to be nonspecific amplification in the majority of the assays, which could be a consequence of the hostile environment offered by feces, resulting in degradation of miRNAs and of other RNA molecules before extraction, yielding short RNA degradation products that might cross-react with some miRNA primers.
Due to accessibility, ease of testing, and extracellular stability of miRNAs in serum, this sample material is an obvious target for biomarker identification.70 The serum miRNA profile could represent any pathological change occurring in an individual dog, and the process of validating serum biomarkers should include ascertaining that changes to serum amounts of a given miRNA can be ascribed to the disease in question. miR-652 could have potential as a serum biomarker for the differentiation between GIC and CIE in dogs, as this was the best performing serum miRNA with regards to the ability to discriminate between GIC and CIE. The relevance of miR-652 is mirrored in the literature, where it has been identified as a plasma biomarker of gastric cancer as well as a prognostic serum biomarker of relapse in colorectal cancer in humans.71, 72 miR-148a-3p amounts were significantly higher in GIC serum relative to both healthy and CIE dogs, which was substantiated by a strong correlation with small RNAseq results. This miRNA has previously been described as a down-regulated serum biomarker of human colorectal cancer relative to healthy controls.73 The discrepancy between these studies could be because of the heterogeneity of both cancer patients and tumor types in our study group. In humans, serum miR-20b is suggested as a biomarker for both gastric cancer and inflammatory bowel disease.74, 75 Whereas our results demonstrated significantly increased amounts of miR-20b in GIC dogs relative to both CIE and healthy dogs, we did not find any significant differences between CIE and healthy dogs for this miRNA. In contrast to human findings, this might suggest that miR-20b could function as a specific GIC serum biomarker in dogs. However, evaluation of miR-20b including a more heterogeneous group of CIE dogs with higher CCECAI scores could serve to clarify the ability of miR-20b in discriminating fully between GIC and CIE dogs.
Our results from the CIE dogs do not support the previous findings of increased amounts of miR-16, miR-122, miR-146a, and miR-147 and decreased amounts of miR-185, miR-192, and miR-223 in serum in dogs with large intestinal inflammatory bowel disease.13 This discrepancy might be because of differences in the investigated dog populations in age, location of disease, as well as differences in CCECAI scores.13
All dogs in this study were systematically characterized through specified inclusion and exclusion criteria, clinical and paraclinical diagnostic workup, scoring, and phenotyping. This is the strength of the current study, and make the results highly applicable when evaluating a dog fulfilling the inclusion criteria. However, it also introduces selection bias to our cohort as many dogs with chronic enteropathy, including food-responsive and antibiotic-responsive enteropathy, as well as all CIE dogs with comorbidities or dogs receiving medication were excluded. We did exclude CIE dogs with primary protein losing enteropathy (PLE) or albumin below 2.0 g/dL. It is unknown if dogs with PLE would have an altered fecal miRNA profile, because of leakage of miRNAs into the intestinal lumen.
As the main objective was to identify dogs with GIC, a subgroup of cancer dogs with comorbidities or that had received medication (or both) were included in the RT-qPCR biomarker identification, but only after statistical evaluation confirming no differences in clinical or miRNA profiles between dogs with comorbidities or that had received medication and dogs that had not. This approach more strongly reflects the dogs encountered in a clinical setting.
There are 3 different tumor types investigated in this study. This serves both as a strength and as a limitation. The number of dogs with each tumor type varies, and reflects the prevalence of diseases, however evaluating our proposed biomarkers on a larger group of dogs with each cancer type could elucidate other associations. Only 7/24 GIC dogs had thoracic imaging to evaluate pulmonary pathology, hence primary tumors or metastases could have been missed.
Another limitation of this study was that all CIE and the majority of GIC dogs had endoscopic pinch biopsies rather than full-thickness biopsies. Though endoscopy is noninvasive and might facilitate localization of lesions, the endoscope is rarely able to access the jejunum, and in some dogs, it can be challenging to gain access to and obtain biopsies from the ileum. Thus, the quality and extent of pinch biopsies are inferior to that of full-thickness biopsies, potentially leading to incorrect diagnoses.76-78 Dogs in all groups had biopsies evaluated by a pathologist at a reference laboratory, however, we did not have a single pathologist review all samples. Even though the interobserver variation between pathologists can be significant,79 histopathologic scoring was not an objective of this study, and was not pursued further.
The CIE dogs in our study had a median CCECAI score of 6 (moderate disease). This CCECAI score was assessed at the time of enrollment and sample collection, and most dogs did have episodes of higher CCECAI scores before or after initial sample collection. Excluding dogs with PLE could also account for the moderate CCECAI score in these population. In future investigations, it will be necessary to address the size and composition of the investigated CIE group, including a larger and more heterogeneous group in validation of the biomarker candidates. This should include other types of chronic enteropathies including food-responsive and antibiotic-responsive enteropathy, PLE, dogs with comorbidities or medical intervention, as well as dogs with GI bleeding. Food-derived miRNAs (xenomiRs) are detectable in feces and blood of humans,80, 81 and it is, therefore, pertinent to evaluate whether meat-rich or raw food diets affect fecal miRNA amounts in dogs.
For the potential future application of the identified GIC miRNA biomarkers in a diagnostic RT-qPCR assay, it is important to consider the performance of the endogenous miRNA normalizers applied in qPCR data analysis.82 Four different miRNAs were employed as normalizers in the present study because of their stability as demonstrated by the NormFinder38 and geNorm39 algorithms. Three were included in these qPCR panels based on either previous reports of suitability as normalizer in feces or serum (miR-16, miR-320),16, 83-86 or based on apparently stable amounts across dog groups in small RNAseq (miR-148b). Additionally, miR-29a was identified as a suitable endogenous normalizer although it was originally included in the panel because of being increased in serum of ulcerative colitis patients.74 This highlights the necessity for the identification of suitable endogenous miRNA qPCR normalizers specifically in the investigated sample material, as good normalizers might not always be found only in literature.
In conclusion, we identified miR-451, miR-223, and miR-27a in feces and miR-20b, miR-148a-3p, miR-652 in serum, as noninvasive biomarker candidates for the optimization of a diagnostic RT-qPCR panel with the potential to differentiate between GIC and CIE in dogs.
ACKNOWLEDGMENT
This study was funded by the Independent Research Fund Denmark - Technology and Production, The Danish Kennel Club, and “Fondet for sygdomsbekæmpelse hos vore familiedyr” (“Fund for Disease Control in our Companion Animals”).
CONFLICT OF INTEREST DECLARATION
The authors declare no conflict of interest.
OFF-LABEL ANTIMICROBIAL DECLARATION
The authors declare no off-label use of antimicrobials.
INSTITUTIONAL ANIMAL CARE AND USE COMMITTEE (IACUC) OR OTHER APPROVAL DECLARATION
The study was approved by the local ethics committees (University of Copenhagen: #2017-9 + 2017-12; Norwegian University of Life Sciences: 14/04723-72, University of Glasgow: Ref 05a/18, University of Edinburgh: VERC#41.18) at all involved institutions as well as by the Animal Experiments Inspectorate under the Ministry of Food, Agriculture, and Fisheries of Denmark, Danish Veterinary and Food Administration (case #2017-15-0201-01353).
HUMAN ETHICS APPROVAL DECLARATION
Authors declare human ethics approval was not needed for this study.
Open Research
DATA AVAILABILITY STATEMENT
All small RNA sequencing data is available from the Sequencing Read Archive (https://www.ncbi.nlm.nih.gov/sra; BioProject ID: PRJNA789413).