New insights into adaptation and population structure of cork oak using genotyping by sequencing
Abstract
Species respond to global climatic changes in a local context. Understanding this process, including its speed and intensity, is paramount due to the pace at which such changes are currently occurring. Tree species are particularly interesting to study in this regard due to their long generation times, sedentarism, and ecological and economic importance. Quercus suber L. is an evergreen forest tree species of the Fagaceae family with an essentially Western Mediterranean distribution. Despite frequent assessments of the species’ evolutionary history, large-scale genetic studies have mostly relied on plastidial markers, whereas nuclear markers have been used on studies with locally focused sampling strategies. In this work, “Genotyping by sequencing” is used to derive 1,996 single nucleotide polymorphism markers to assess the species’ evolutionary history from a nuclear DNA perspective, gain insights into how local adaptation is shaping the species’ genetic background, and to forecast how Q. suber may respond to global climatic changes from a genetic perspective. Results reveal (a) an essentially unstructured species, where (b) a balance between gene flow and local adaptation keeps the species’ gene pool somewhat homogeneous across its distribution, but still allowing (c) variation clines for the individuals to cope with local conditions. “Risk of Non-Adaptedness” (RONA) analyses suggest that for the considered variables and most sampled locations, (d) the cork oak should not require large shifts in allele frequencies to survive the predicted climatic changes. Future directions include integrating these results with ecological niche modeling perspectives, improving the RONA methodology, and expanding its use to other species. With the implementation presented in this work, the RONA can now also be easily assessed for other organisms.
1 INTRODUCTION
Understanding how and at which rate species respond to global climatic change in their environmental context is becoming an increasingly important question due to the pace at which these are taking place (Kremer et al., 2012; Primack et al., 2009). To avoid obliteration, species may respond to such changes by either altering their distribution range, or by adapting to the new conditions. The latter can occur “instantly,” due to phenotypic plasticity, or across several generations, by local adaptation (Aitken, Yeaman, Holliday, Wang, & Curtis-McLane, 2008). The kind of response species can provide is known to depend on factors such as location, distribution range, and/or genetic background (Gienapp, Teplitsky, Alho, Mills, & Merilä, 2008; Ohlemuller, Gritti, Sykes, & Thomas, 2006).
Tree species are characterized by sedentarism and long lifespan and generation times, allied with generally large distribution ranges and capacity for long-distance dispersal through pollen and seeds (Kremer et al., 2012). These traits make them interesting subjects to study regarding their response to global climatic changes (Thuiller et al., 2008).
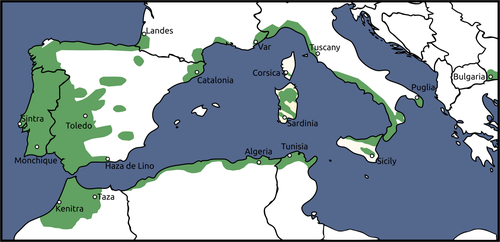
In this work, we address the case of the cork oak (Quercus suber L.). With a distribution ranging most of the West Mediterranean region (Figure 1), this oak species is the most selective evergreen oak of the Mediterranean basin in terms of precipitation and temperature conditions (Vessella, López-Tirado, Simeone, Schirone, & Hidalgo, 2017). European oaks, in particular, are known to have endured past climatic alterations, but how they can cope with the current, rapidly occurring changes is not yet fully understood (Kremer et al., 2012; Kremer, Potts, & Delzon, 2014). Despite this tree’s ecological and economic importance, there is yet much to learn regarding the consequences of global climatic change on its future (Benito Garzón, Sánchez de Dios, & Sainz Ollero, 2008).
Some recent works have attempted to answer this very question, but focusing on range expansion and contraction with the assumption of a genetically homogeneous species and niche conservationism (Correia, Bugalho, Franco, & Palmeirim, 2017; Vessella et al., 2017). Both these studies also highlight the need for a genetic study regarding the adaptation potential of Q. suber. Unlike what happen in other oak species (Rellstab et al., 2016), studies integrating genetic information and response to climatic alterations of Q. suber (e.g. (Modesto et al., 2014)) are rare and of small scale (Ramírez-Valiente, Valladares, Huertas, Granados, & Aranda, 2011). Even though this study made the important assessement that some cork oak traits can be associated with genetic variants, its local geographic scope, combined with the relatively low number if used markers, limits its utility in a distribution wide perspective. Large-scale information regarding Q. suber's gene flow patterns and local adaptation dynamics is paramount to understanding the species’ potential to endure rapid climatic changes through adaptation (Savolainen, Lascoux, & Merilä, 2013).
In general terms, to predict a species’ response to change (Kremer et al., 2012), it is fundamental to know both its genetic architecture of adaptive traits (Alberto et al., 2013) and its evolutionary history (Kremer et al., 2014). However, the very nature of genetic and genomic data hampers the distinction of selection signals from other processes (McVean & Spencer, 2006), especially demographic events (Bazin, Dawson, & Beaumont, 2010). In order to disentangle population structure (mostly shaped by gene flow, inbreeding, and genetic drift) and selection (Foll, Gaggiotti, Daub, Vatsiou, & Excoffier, 2014), recent methods incorporate population structure information to detect adaptation (Gautier, 2015; Günther & Coop, 2013). Likewise, methods to accurately estimate population structure should be performed without loci known to be under selection (De Kort et al., 2014).
In nonmodel organisms such as the cork oak, loci of adaptive value can potentially be identified by two kinds of methods—outlier analyses and environmental association analyses (EAA). While the former identify loci that depart from the expected allele frequencies as under selection (Foll & Gaggiotti, 2008; Vitalis, Gautier, Dawson, & Beaumont, 2014), they do not indicate what which loci are responding to (Gautier, 2015). The latter, while being able to associate the markers to an external covariate, are limited to detecting linear relations, and cannot assert whether or not the identified correlations are of causative nature (Gautier, 2015).
The evolutionary history of Q. suber has been studied in the past using multiple methodologies and in different geographic ranges. The most recent large-scale studies on the subject suggest that cork oak is divided into four strictly defined lineages (Cosimo et al., 2009; Magri et al., 2007). Two of these lineages range from the south-east of France, to Morocco, including the Iberian Peninsula and the Balearic Islands, a third lineage ranges from the Monaco region to Algeria and Tunisia, including the islands of Corsica and Sardinia. The fourth lineage spans the entire Italic peninsula, including Sicilia. Based only on plastidial markers, these lineages have been shown to hardly share any haplotypes (Magri et al., 2007). Notwithstanding, later works based on nuclear DNA have hinted at a different scenario, where the species is not as strictly divided (Costa et al., 2011; Ramírez-Valiente, Valladares, & Aranda, 2014). These works are, however, limited in either geographic scope or number of markers to confidently conclude that such segregation is only present in plastidial markers.
Genomic resources represent a new way to study the genetic mechanisms responsible for local adaptation (Rellstab, Gugerli, Eckert, Hancock, & Holderegger, 2015) through the use of EAA, which correlate environmental data with genetic markers, thus highlighting loci putatively involved in the adaptation process (Rellstab et al., 2016). The same methods, can thus, in principle, be used to assess the degree of maladaptation to predicted future local conditions (Rellstab et al., 2016). The risk of non-adaptedness (RONA) method was developed with this very goal (Rellstab et al., 2016). In short, for every significant association between a single nucleotide polymorphism (SNP) and an environmental variable, the RONA method plots each location's individuals’ allele frequencies vs. the respective environmental variable. This is done for both the current value and future prediction. A correlation between allele frequencies and the current variable values is then calculated and the corresponding best fit line is inferred. The distance between the fitted line and the two coordinates is then compared per location and its normalized difference is considered the RONA value for each association and location (which can vary between 0 and 1). In theory, the higher the difference in conditions between the current values and the prediction, the more the studied species should have to shift its allele frequencies to survive in the location under the new conditions. Despite the innovation and importance of the method for the general scientific community, in the original paper, RONA is applied only for the work's case study (calculating RONA values for several Swiss species of Quercus based on candidate genes), and no public implementation is provided. Applying this kind of methodology to Q. suber would fill the gap mentioned in (Correia et al., 2017; Vessella et al., 2017), that multidisciplinary approaches are required to more accurately provide sound recommendations for the conservation of forests.
In the present work, a panel of SNP markers derived from the Genotyping by sequencing (GBS) technique (Elshire et al., 2011) was developed to accomplish the following goals: (a) attempt to infer the species’ genetic structure and evolutionary history, (b) detect signatures of natural selection, and (c) investigate the adaptation potential of Q. suber based on the RONA method developed and presented on (Rellstab et al., 2016).
2 MATERIALS AND METHODS
2.1 Sample and environmental data collection
In order to provide a comprehensive view of the species genetic background, samples were collected from 17 locations spanning most of Q. suber's distribution. Fresh leaves were collected from six individuals from, Bulgaria, Corsica, Kenitra, Monchique, Puglia, Sardinia, Sicilia, Tuscany, Tunisia, and Var, and from five individuals from Algeria, Catalonia, Haza de Lino, Landes, Sintra, Taza, and Toledo for a total of 95 individuals (Table 1, Figure 1). It is worth noting that trees from Bulgaria are not of natural origin, but rather the result of human introduction from Iberian locations (Borelli & Varela, 2000; Petrov & Genov, 2004).
| Sample site | Latitude (decimal deg.) | Longitude (decimal deg.) | Number of sampled individuals |
|---|---|---|---|
| Algeria | 36.5400 | 7.1500 | 5 |
| Bulgaria | 41.43 | 23.17 | 6 |
| Catalonia | 41.8500 | 2.5333 | 5 |
| Corsica | 41.6167 | 8.9667 | 6 |
| Haza de Lino | 36.8333 | −3.3000 | 5 |
| Kenitra | 34.0833 | −6.5833 | 6 |
| Landes | 43.7500 | −1.3333 | 5 |
| Monchique | 37.3167 | −8.5667 | 6 |
| Puglia | 40.5667 | 17.6667 | 6 |
| Sardinia | 39.0833 | 8.8500 | 6 |
| Sicilia | 37.1167 | 14.5000 | 6 |
| Sintra | 38.7500 | −9.4167 | 5 |
| Taza | 34.2000 | −4.2500 | 5 |
| Toledo | 39.3667 | −5.3500 | 5 |
| Tunisia | 36.9500 | 8.8500 | 6 |
| Tuscany | 42.4167 | 11.9500 | 6 |
| Var | 43.1333 | 6.2500 | 6 |
| Total | — | — | 95 |
Most samples were collected from an international provenance trial (FAIR I CT 95 0202) established at “Monte Fava,” Alentejo, Portugal (38°00′ N; 8°7′ W) (Varela, 2000), except Portuguese and Bulgarian samples, which were collected directly from their native locations. The collected plant material was stored at −80°C until DNA extraction.
Altitude, latitude, and longitude spatial variables (Varela, 2000) were recorded for each of the native sampling sites. Nineteen Bioclimatic (BIO) variables, BIO1 to BIO19, were collected from the WorldClim database (Hijmans, Cameron, Parra, Jones, & Jarvis, 2005) at 30 arc-seconds (~1 km) resolution for both “Current conditions ~1960–1990” and “Future” predictions for 2070, using two different Representative Concentration Pathways (RCPs), rcp26 and rcp85 for the following “Global Climate Models” (GCMs): BCC-CSM1–1, CCSM4, GFDL-CM3, GISS-E2-R, HadGEM2-ES, IPSL-CM5A-LR, MRI-CGCM3, MPI-ESM-LR, and NorESM1-M (IPCC, 2014) as these are available under permissive licenses and calculated for both rcp26 and rcp85. Instead of using the GCMs directly, an average of the values was obtained for each coordinate, and merged into a single dataset, for both used RCPs (Tables S1 and S2, respectively). Data were extracted from the GeoTiff files using a python script, layer_data_extractor.py (https://github.com/StuntsPT/Misc_GIS_scripts) as of commit “bd36320”.
Correlations between present Bioclimatic variables were assessed using Pearson's correlation coefficient as implemented in the R script eliminate_correlated_variables.R (https://github.com/JulianBaur/R-scripts) as of commit “43e6553,” which resulted in the exclusion of six variables due to high correlation (r > 0.95). Each sampling location was thus characterized by three spatial variables and 13 environmental variables (Table S3).
2.2 Library preparation and sequencing
Genomic DNA was extracted from liquid nitrogen grounded leaves of all samples collected for this work using the kit “innuPREP Plant DNA Kit” (Analytik Jena AG), according to the manufacturer's protocol.
The total amount of extracted DNA was quantified by spectrophotometry using a Nanodrop 1000 (Thermo Scientific) and integrity verified on Agarose gel (0.8%). DNA samples were then diluted to a concentration of ~100 ng/μl and plated for genotyping.
DNA samples were then outsourced to “Genomic Diversity Facility,” at Cornell University” for genotyping using the “Genotyping by sequencing” (GBS) technique as described in (Elshire et al., 2011). Samples were shipped in a single 96-well plate with one “blank” well for negative control. Sequencing was performed according to the standard protocol on a single Illumina HiSeq 2000 flowcell using the low-frequency cutter enzyme “EcoT22I,” due to the large size of Q. suber's genome.
2.3 Genomic data analyses
The raw GBS data were analyzed using the program ipyrad v0.7.24, which is based on pyrad (Eaton, 2014), using an “anaconda” environment containing—muscle v3.8.31 (Edgar, 2004) and vsearch v2.7.0 (Rognes, Flouri, Nichols, Quince, & Mahé, 2016). A de novo sequence assembly was performed, but mtDNA and cpDNA reads were “baited” out by ipyrad's mode “denovo-reference” using the complete mitochondiral genomes of Populus davidiana (KY216145.1) (Choi et al., 2017), Pyrus pyrifolia (KY563267.1) (Chung, Lee, Kim, & Kim, 2017) and Rosa chinensis (CM009589.1) (Raymond et al., 2018), and chloroplastidial genomes of Quercus rubra (JX970937.1) (Alexander & Woeste, 2014), Quercus aliena (KU240007.1) and Quercus variabilis (KU240009.1) (Yang et al., 2016). This ensured that mtDNA and cpDNA reads were filtered from downstream analyses. Parameters included GBS as datatype, clustering threshold of 0.85, mindepth of 8 and maximum barcode mismatch of 0. Each sampling site had to be represented by at least three individuals for a SNP to be called, except the locations of Kenitra and Taza, where only one individual was required due to the lower representation of these sampling sites. Full parameters can be found in Datafile S1. The demultiplexed “fastq” files were submitted to NCBI's Sequence Read Archive (SRA) as “BioProject” PRJNA413625.
Downstream analyses were automated using “GNU Make.” This file, containing every detail of every step of the analyses for easier reproducibility, can be found in gitlab (https://gitlab.com/StuntsPT/Qsuber_GBS_data_analyses, tag “v03”). For improved reproducibility, a docker image with all the software, configuration files, parameters and the Makefile, ready to use is also provided (https://hub.docker.com/r/stunts/q.suber_gbs_data_analyses/, tag “v03”). The intent is not to allow the analyses process to be treated as a “black box,” but rather to provide a full environment that can be reproduced, studied, and modified by the scientific community.
Processed data from ipyrad were then filtered using VCFtools v0.1.14 (Danecek et al., 2011) with the following criteria: Each sample has to be represented in at least 40% of the SNPs, and after this, each SNP has to be represented in at least 80% of the individuals. Furthermore, due to the relatively small sample size, the minimum allele frequency (MAF) of each SNP has to be at least 0.03 for it to be retained.
In order to minimize the effects of linkage disequilibrium, downstream analyses were performed using only one SNP per locus, by discarding all but the SNP closest to the center of the sequence in each locus. This sub-dataset was obtained using the python script vcf_parser.py (https://github.com/CoBiG2/RAD_Tools/blob/master/vcf_parser.py) as of commit “0893296”.
All file format conversions were performed using PGDSpider v2.1.0.0 (Lischer & Excoffier, 2012), except for the BayPass and SelEstim formats, where the scripts geste2baypass.py (https://github.com/CoBiG2/RAD_Tools/blob/master/geste2baypass.py) and gest2selestim.sh (https://github.com/Telpidus/omics_tools) as of commit “b99636e” and “f74f66b,” respectively, were used, since the used version of PGDSpider does not handle either of these formats.
Descriptive statistics, such as Hardy–Weinberg Equilibrium (HWE), FST and FIS were calculated using Genepop v4.6 (Rousset, 2008). The same software was further used to perform Mantel tests to determine an eventual effect of Isolation by Distance (IBD) by correlating “'F/(1 − F)'-like with common denominator” with “Ln(distance)” following on 1,000,000 permutations. This test was performed excluding individuals sampled from Bulgaria due to their introduced origin.
2.4 Outlier detection and environmental associations
Outlier detection was performed using two programs: SelEstim v1.1.4 (Vitalis et al., 2014) (50 pilot runs of length 1,000 followed by a main run of length 10⁶, with a burnin of 1,000, a thinning interval of 20, and a detection threshold of 0.01) and BayeScan v2.1 (Foll & Gaggiotti, 2008) (20 pilot runs of length 5,000 followed by a main run of 500,000 iterations, a burnin of 50,000, a thinning interval of 10, and a detection threshold of 0.05) (full commands and parameters are available in Datafile S2), since these methods show the lowest rate of false positives (Narum & Hess, 2011; Vitalis et al., 2014). Only SNPs indicated as outliers by both programs were considered outliers for the purpose of this work. This was done to further reduce the chance of false positives, which is a known issue in this type of analyses (Gautier, 2015; Vitalis et al., 2014).
The software BayPass v2.1 (Gautier, 2015) wrapped under the script Baypass_workflow.R (https://gitlab.com/StuntsPT/pyRona/blob/master/pyRona/R/Baypass_workflow.R) from pyRona v0.1.3 was used to assess associations of SNPs to environmental variables using the “AUX” model (20 pilot runs of length 1,000, followed by a main run of length 500,000 with a burnin of 5,000 and a thinning interval of 25). Any association with a Bayes Factor (BF) above 15 was considered significant. Association analyses were performed excluding individuals from Bulgaria sampling site for the same reasons as in the Mantel tests.
Sequences containing outlier loci or SNPs associated to an environmental variable were queried against the genome of Q. lobata (Sork et al., 2016) v1.0 using blast v2.2.28+ (Altschul et al., 1997) with an e-value threshold of 0.00001.
2.5 Population structure
Two distinct methods were used for clustering the individuals in order to understand the general pattern of individual or population grouping, namely, principal components analysis (PCA) and MavericK (Verity & Nichols, 2016), which is based on structure (Pritchard, Stephens, & Donnelly, 2000).
The PCA was performed with snp_pca_static.R (https://github.com/CoBiG2/RAD_Tools/blob/master/snp_pca_static.R) as of commit “bb2fc45”.
In order to correctly interpret clustering analyses results, it is important to estimate the value of K, which represents how many demes the data can be clustered into. The software MavericK is especially interesting for cluster estimation due to its innovative method for estimating K, called “Thermodynamic Integration” (TI), which has shown superior performance in this task relative to other methods (Verity & Nichols, 2016). Analysis was divided into two stages: an initial single “pilot” stage which ran for 5,000 iterations, with a burnin of 500 using an admixture model, a free alpha parameter of “1” and “thermodynamic integration” (TI) turned off. This stage was used to infer tuned alpha and alphaPropSD values which were used in the subsequent “tuned” stage as parameters for the admixture model. This stage was comprised of five runs of 10,000 iterations (10% burnin), with TI turned on and set to 20 rungs of 10,000 samples with 20% burnin. MavericK was wrapped under Structure_threader v 1.2.2 (Pina-Martins, Silva, Fino, & Paulo, 2017) and was run for values of K between 1 and 8. The most suitable value of K was estimated using the TI method. Full parameter files are available as Datafile S2.
The same methodology was used on two more datasets derived from the original data. On one, only SNPs considered “neutral” were used, in order to obtain an unbiased population structure (De Kort et al., 2014). On the other one, only SNPs considered “non-neutral” were used, which should not be interpreted as population structure, but rather as an indication of whether local adaptation is responsible for the observed pattern.
2.6 Risk of non-adaptedness
The software pyRona was developed in this work as the first public implementation of the method described in (Rellstab et al., 2016) called “Risk of non-adaptedness” (RONA). This method provides a way to represent the theoretical average change in allele frequency at loci associated with environmental variables required for any given population to cope with changes in that variable. The program source code is hosted on public repositories, under a GPLv3 license, and can be downloaded free of charge at https://gitlab.com/StuntsPT/pyRona.
pyRona has a complete user manual, with installation instructions, usage patterns, and a graphical method description.
The RONA method as implemented in pyRona, however, is slightly different from the original method description (Datafile S3). Namely, instead of ranking environmental factors by p-value of the difference test between present and future values like the original description, pyRona will rank the environmental factors by the number of associations. Furthermore, the average RONA value provided by pyRona is weighted by the R2 value of each involved correlation, unlike the original, which uses unweighted means.
In this work, two alternative climate prediction models were used to calculate a RONA value for each location in pyRona v0.1.3: a low emission scenario (RCP26) and a high emission scenario (RCP85) (IPCC, 2014) in order to account for uncertainties in the models’ assumptions. Any associations flagged by Baypass with a BF above 15 were considered relevant and included in the RONA analysis. The three nongeospatial environmental variables most frequently associated with SNPs were selected for determining generic RONA values.
3 RESULTS
Genotyping by sequencing (Elshire et al., 2011), a technique based on restriction enzyme genomic complexity reduction followed by short-read sequencing, was employed to discover SNP markers from a total of 95 Q. suber individuals sampled from 17 geographical locations (Table 1).
A total of 225,214,094 reads (100 bp) generated by the GBS assay was processed by ipyrad (Eaton, 2014) computational pipeline. The first analytical step consisted in the assembly of raw reads into 4,548 distinct contiguous sequence fragments (genomic loci), from which an initial set of 8,978 SNPs were flagged. Twelve Q. suber samples were discarded due to low sequence representation during the assembly process, resulting in the retention of 83 individuals. After filtering according to the criteria presented in the Methods section 2.3, 1,996 SNPs remained, which were used for all further analyses. This filtering process additionally removed two samples which were not represented for more than 55% of the markers, and therefore, only 81 samples were used in the analyses (Table S4).
The calculated FIS values for each sampling site are available in Table S4. These range from −0.0262 (Var) to 0.1145 (Puglia) with an average value of 0.0666. Pairwise FST values are available in Table S5. These range from 0.0028 between Sardinia and Tuscany to 0.1216 between Landes and Var (average FST of 0.0541).
When looking at HWE results per marker, of the 1,996 SNPs, 172 (~9%) reveal a heterozygote deficit, whereas 88 (~4%) reveal a deficit of homozygotes. Individual sampling sites are comprised of two few individuals to achieve biologically meaningful results. The performed Mantel test revealed no evidence of IBD among Q. suber individuals.
3.1 Outlier detection and environmental association
Population differentiation and ecological association approaches (François, Martins, Caye, & Schoville, 2016) were employed aiming at the identification of loci targeted by selection. In the first strategy, highly differentiated loci among populations, measured as outliers in FST distribution, were detected by the software BayeScan and SelEstim uncovering 29 and 17 outlier SNPs, respectively (Table S6). All of the loci considered under outliers by SelEstim were also present in the set of loci flagged as outlier by BayeScan. This set of 17 common markers was considered as being putatively under the effect of natural selection.
For a functional characterization of these loci, the draft genome sequence of Q. lobata was used as a proxy for similarity searches. None of the 17 sequences revealed significant matches to Q. lobata's genome scaffolds.
The ecological association approach was carried out using the software BayPass and yielded 274 associations between 249 SNPs and 12 of the 16 tested environmental variables (no associations were found with “Altitude,” “Temperature Annual Range,” “Precipitation of Wettest Month,” or “Precipitation Seasonality”). These associations can be found in Table S7. Despite this relatively high number of associations, it is important to note that 70 of these associations were between a SNP and a geospatial variable: 12 associations with “Latitude” and 58 with “Longitude.” Of all environmental variables, the one with most markers associated is “Precipitation of Driest Month” with 71 associations, followed by “Isothermality” with 35 associations, and “Mean Temperature of Driest Quarter” with 29 associations.
Sequences containing 22 of the 249 markers associated with environmental variables were matched to entries in the Q. lobata genome; however, of these, only 10 were annotated (Table 2).
| SNP name | Note (Similar to) | Associations |
|---|---|---|
| SNP 158 | TRE1: Trehalase (Arabidopsis thaliana) | Mean Temperature of Driest Quarter |
| SNP 168 | PER47: Peroxidase 47 (Arabidopsis thaliana) | Precipitation of Driest Month |
| SNP 233 | CPSF160: Cleavage and polyadenylation specificity factor subunit 1 (Arabidopsis thaliana) | Annual Mean Temperature |
| SNP 333 | Ascc1: Activating signal cointegrator 1 complex subunit 1 (Mus musculus) | Mean Temperature of Driest Quarter |
| SNP 455 | GLCAT14A: Beta-glucuronosyltransferase GlcAT14A (Arabidopsis thaliana) | Precipitation of Driest Month |
| SNP 619 | GBP6: Guanylate-binding protein 6 (Pongo abelii) | Precipitation of Driest Month |
| SNP 834 | NAC098: Protein CUP-SHAPED COTYLEDON 2 (Arabidopsis thaliana) | Longitude |
| SNP 880 | TPP1: Thylakoidal processing peptidase 1%2C chloroplastic (Arabidopsis thaliana) | Mean Temperature of Warmest Quarter |
| SNP 1134 | EMB2654: Pentatricopeptide repeat-containing protein At2g41720 (Arabidopsis thaliana) | Mean Temperature of Driest Quarter |
| SNP 1589 | At1g19525: Pentatricopeptide repeat-containing protein At1g19525 (Arabidopsis thaliana) | Temperature Seasonality |
The union of the outlier loci set and the set of loci associated with at least one environmental variable resulted in a dataset of 259 SNPs which were deemed “non-neutral” (seven SNPs were common to both loci sets). The remaining 1737 SNPs were grouped in another sub-dataset, deemed “neutral.”.
3.2 Population structure
Clustering analyses were used to infer the current population structure of Q. suber in the West Mediterranean. The TI method implemented in the software MavericK determined the best K value to be “1” on all datasets. Despite this assessment, the presented plots are always with K = 2 (Figure 2), but with strong evidence that the data do not support structuring of any kind. Q-plots for values of K above 2 were always either reduced to two clusters, or to every individual being roughly equally divided into fractions of all clusters (Figure S1).
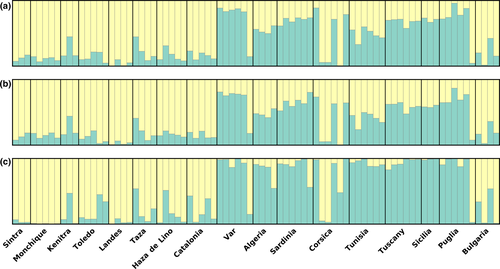
The Q-matrix plot showing the relatedness of each genotype to each considered deme of MavericK’s results produced using all loci (Figure 2a) can be interpreted as a rough split between western individuals (from locations Sintra, Monchique, Kenitra, Toledo, Landes, Taza, Haza de Lino, and Catalonia), which are mostly, but not completely, assigned to cluster “1” and eastern ones (from locations Var, Algeria, Sardinia, Corsica, Tunisia, Tuscany, Sicilia and Puglia), which are mostly assigned to cluster “2”. Individuals from Bulgaria are a notable exception, since individual genotypes are mostly assigned to cluster “1” similar to those of individuals from western locations, likely due to the species’ introduced origin (Varela, 2000). However, this West–East split is somewhat fuzzy, as individuals’ genomes are never completely attributed to a single cluster. In fact, most individuals have a considerable part of their genome attributed to both cluster “1” and “2.” Furthermore, individuals from some eastern locations have their genomes almost completely attributed to cluster “1” (Var 21, Corsica 3, Corsica 11, Corsica 14, and Puglia 5), and all individuals from Tunisia and Algeria are almost equally split between both clusters.
The Q-plot obtained using the “neutral” loci subset (Figure 2b) is nearly identical to the one with all the loci, but with individual genomes from eastern locations being slightly more assigned to cluster “1” than in Figure 2a, and can be interpreted in the same way.
The Q-plot produced using only the 259 (12.9%) “non-neutral” loci (Figure 2c), however, does bear a different clustering pattern from the previous ones. In this case, the East–West split is more evident, as eastern individual genomes’ attribution to each cluster is not as evenly split, but rather displays a more pronounced attribution to cluster “2” than in Figure 2a. The opposite is also true for western individuals, but to a lesser extent.
The PCA clustering method (largest eigenvector values of 0.0405 and 0.0299) is essentially concordant with the previous methods, revealing two loosely defined groupings (Figure S2).
3.3 Risk of non-adaptedness (RONA)
A summary of the RONA analyses for both low (RCP26) and a high (RCP85) emission scenario predictions can be found in Figure 3 and Table S8. The most represented environmental variables are “Precipitation of Driest Month” (71 SNPs, mean R2 = 0.1570), “Isothermality” (35 SNPs, mean R2 = 0.2143), and “Mean Temperature of Driest Quarter” (29 SNPs, mean R2 = 0.1501). The values of RONA per sampling site are always higher for RCP85 than for RCP26, except for “Precipitation of Driest Month” in Tunisia where RCP85 has a lower RONA than RCP26, and in Kenitra where they are the same (the “Precipitation of Driest Month” variable in Kenitra is not predicted to change from current conditions of 0 mm2 regardless of the model).
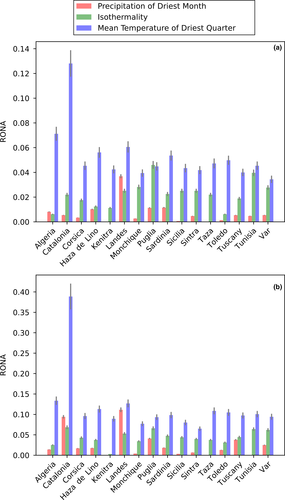
Under the RCP26 predictions, the highest RONA values for “Precipitation of Driest Month” are Landes (0.0369), for “Isothermality” is Puglia (0.0461), and for “Mean Temperature of Driest Quarter” is Catalonia (0.1281). Under the RCP85 predictions, Landes presents the highest RONA for “Precipitation of Driest Month” (0.1115) and Catalonia presents the highest values of RONA for “Mean Temperature of Driest Quarter” (0.3888) and “Isothermality” 0.0686). It is important to note that the high RONA values of Catalonia are approximately twice as high as the second highest RONA value on the RCP26 prediction and close to three times as high for RCP85, marking this location as the most likely to become deprived of cork oak individuals in the future.
4 DISCUSSION
In this study, Q. suber individuals were sampled across the species’ distribution range to assess population structure, impact of local adaptation, and provide an estimate of the RONA value of each sampled location.
Due to the relatively large size of Q. suber's genome (Zoldos, Papes, Brown, Panaud, & Siljak-Yakovlev, 1998) a genome reduction technique, GBS, was used to discover SNPs for this species. There is no “standard” parameter set to call SNPs on GBS datasets, since this will ultimately depend on the organism being studied. The stringent approach used in this study was, however, deemed preferable to alternatives that could result in more SNPs being called at the cost of lowering confidence in the called variants, eventually biasing analyses results. In fact, since no biological replicates were performed for this study, a conservative approach was always preferred as to minimize biases in the results.
After stringent quality filtering, a set of 1,996 SNPs was used in this study. This number is lower than that of some studies with similar data (Berthouly-Salazar et al., 2016), which obtained ~22 k SNPs (albeit using a more frequent cutting enzyme), but still more than (De Kort et al., 2014), which obtained 1630 SNPs, very close to that of (Escudero, Eaton, Hahn, & Hipp, 2014) and (Pais, Whetten, & Xiang, 2017). Even though this number may seem small, in the universe of Q. suber's genome of ~750 Mbp, this is to date the largest number of molecular markers available for this species and represents a step forward to increase the power of population genetics studies.
4.1 Population genetic structure
Past studies (Magri et al., 2007) have characterized Q. suber as a highly structured species, with an evolutionary history shaped by large effect events, such as plate tectonics. These were, however, mostly based on plastidial DNA data, which are known to not always provide a comprehensive view on a species’ evolutionary history (Kirk & Freeland, 2011). The nuclear markers developed for this work provide a somewhat different perspective.
Hardy–Weinberg Equilibrium analysis revealed that few individual markers deviated from expectations. Only ~9% reveal a heterozygote deficit, and only ~4% reveal a deficit of homozygotes. These values do not indicate the presence of assembly bias.
The obtained values of FIS are higher than those of unstructured European oaks when analyzed with the same type of markers, such as Quercus robur or Quercus petraea (Guichoux et al., 2013), but are nonetheless relatively low in general, which is compatible with low levels of population structuring.
Similar to what is observed with FIS, FST values are on average (0.0541) higher than on the above-mentioned unstructured oak species (0.0125) (Guichoux et al., 2013), but lower than other well-structured trees such as eucalypti (0.095) (Cappa et al., 2013). These results corroborate what the clustering analyses reveal: an incomplete segregation of the species in two clusters, as seen on Figure 2. Although clustering analyses using all loci do not provide a clear structuring signal (and the “TI” method clearly favors a scenario of a single large panmictic population), the produced Q. suber Q-plots do show some degree of segregation between western and eastern individuals. This can be derived both from Figure 2a,b, which are very similar and can be interpreted in the same way—as incomplete segregation between individuals from eastern and western locations.
Figure 2c, where the Q-plot was produced using only loci putatively under selection, should not be used to infer population structure, but can be compared to the Q-plot obtained using only “neutral” loci to interpret the role of local adaptation in shaping Q. suber's genetic background. In Figure 2c, the division between western and eastern individuals is clearer than in Figure 2a,b. Furthermore, the generally observed difference pattern is similar to what can be seen in the locations of “Monchique” and “Sardinia”: Individual attributions to the “dominant” cluster in the “neutral” Q-plot become even more pronounced in the “non-neutral” Q-plot. This is expected if local adaptation is responsible for these differences (otherwise, the differences between “neutral” and “non-neutral” Q-plots should be more random). This evidence, combined with the relatively low pairwise FST and FIS values, suggests a balance between local adaptation and gene flow. Whereas the former is responsible for maintaining the species’ standing genetic variation across the species range and the latter for the species’ response to local environmental differences. Intense gene flow would also explain the relatively low proportion of outlier SNPs, which may be counteracting reactions to weak selective pressures. At the same time, this balance may provide the species with a relatively large genetic variability to respond to strong selection (De Kort et al., 2014; Kremer et al., 2012).
Data from this work do not seem to support the four lineages hypothesis proposed in (Magri et al., 2007); however, it is also not incompatible with it, if it is assumed that nuDNA and cpDNA can have different evolutionary histories. In fact, it has been argued that for other tree species plastidial lineages exist due to population contractions and expansions from glacial refugia, but high gene flow erases any evidence of their existence in the nuclear genome (Eidesen et al., 2007).
- Balance between gene flow and local adaptation is responsible for both creating and maintaining the current level of nuclear divergence. Whereas local adaptation tends to cause divergence between contrasting regions, this effect is countered by species wide gene flow. Population contractions in refugia locations during glacial periods explain the occurrence of plastidial lineages, which are absent in the nuclear genome due to very intense gene flow.
- Differential hybridization of Q. suber with Q. cerris in the East (Bagnoli et al., 2016) and with Q. ilex s.l. in the West (Burgarella et al., 2009) is responsible for the observed nuDNA structuring pattern and balance between gene flow and local adaptation is responsible for maintaining it. Combination of these phenomena can thus be considered the cause for the observed levels of East–West differentiation. Since Q. suber always acts as a pollen donor in these hybridization events (Boavida, Silva, & Feijó, 2001). Under this hypothesis, Q. suber would maintain a high nuclear population effective, even during glacial periods, but restrict plastidial lineages’ geographic scope as suggested in (López de Heredia, Carrión, Jiménez, Collada, & Gil, 2007), which is further supported by the different dispersal capabilities of pollen and acorns (Sork, 1984). This scenario would result in large effective population size differences between nuDNA and cpDNA, which can be an alternative explanation for cpDNA lineages to simple population contractions to glacial refugia.
The proposed hypotheses are supported by the SNP data presented here, but further studies are needed to confirm them. As such, the issue will remain open for investigation.
4.2 Outlier detection and EAA
The method used to detect outlier loci flagged ~0.9% of the total SNPs, which is in line with what was found on other similar studies (Berdan, Mazzoni, Waurick, Roehr, & Mayer, 2015; Chen et al., 2012). Of the 17 outlier markers found, none could be matched to an annotated location in Q. lobata's genome. This is likely due to a combination of factors, such as the distance between Q. suber and Q. lobata, and the incomplete annotation of Q. lobata's genome. On the other hand, it emphasizes the need for more genomic resources in this area, which can potentially provide important functional information of these SNPs in Q. suber's genome, which will at least for now remain unknown.
The EAA served two purposes in this work. On one hand, the reported associations work as a proxy for detecting local adaptation, and on the other hand, allow the attribution of a RONA score to each sampling site. Q. suber is known to be very sensitive to precipitation and temperature conditions (Vessella et al., 2017), and as such, it was expected beforehand that some of the markers obtained in this study were to be associated with some of these conditions (Rellstab et al., 2016). In order to understand how important the found associations are for the local adaptation process, it is necessary to understand the putative function of the genomic region where each SNP was found. Querying the available sequences against Q. lobata's genome annotations has provided insights regarding some of the markers’ sequences putative function. The proportion of sequences that were a match to an annotated region, however, is rather small—only ~4.4% of the queried sequences could be matched to such regions.
Of the 10 SNPs associated with an environmental variable that returned hits to annotated regions of Q. lobata's genome, two were matched to regions annotated as close to animal genes, and one matched a region annotated as a chloroplastidial region, leaving seven SNPs as interesting to explore for downstream analyses. While all these associations are potentially interesting to explore, doing so falls outside the grander scope of this work.
Of these markers, it is interesting to remark, that SNP 158, associated with the variable “Mean Temperature of Driest Quarter”, for example, is located in a region annotated as “Similar to TRE1: Trehalase,” which is known to play a role in drought stress (Houtte et al., 2013). Likewise, SNP 168, associated with the variable “Precipitation of Driest Month,” is located in a region matching the annotation of “Similar to PER47: Peroxidase 47,” which is known to play a role in drought response (Li et al., 2017).
Like these two examples, more of the SNPs found have associations to environmental variables which are putatively located in genes involved in functions which are important in responding to the very variables they are associated with. This fact flags these markers as particularly useful to focus on in downstream studies.
4.3 Risk of non-adaptedness
Although the RONA method is a greatly simplified model (its limitations are described in Rellstab et al., 2016), it provides an initial estimate of how affected Q. suber is likely to be by environmental changes (at least as far as the tested variables are concerned). Furthermore, it is important to remark that due to Baypass being limited to a univariate method, the same constraint also applies to the RONA analysis, meaning that multiloc associations are not considered.
The implementation developed for this work named pyRona suffers from most of the same limitations as the original application, even though it is based on an arguably superior association detection method (Gautier, 2015), (although the original LFMM (Frichot, Schoville, Bouchard, & François, 2013) method is also available to use in pyRona since version 0.3.0) and introduces a correction to the average values based on the R2 of each marker association by using weighted means. The automation achieved by using this new implementation easily allows two different emission scenarios (RCP26 and RCP85) to be tested and compared.
With the exception of Catalonia, which seems to have an exceptionally high highest RONA value under both prediction models, the other locations present relatively low RONA values for the tested variables. The variable “Mean Temperature of Driest Quarter” appears to be the tested variable that requires the greatest changes in allele frequencies to ensure adaptation of the species to the local projected changes. These RONA values are nevertheless smaller than those presented in (Rellstab et al., 2016). This might be due to various factors, such as the different variables tested, the geographic scope of the study, the species’ respective tolerance to environmental ranges, the differences between species’ standing genetic variation, the association detection method, or, more likely, a combination of several of these factors.
Notwithstanding, the obtained results seem to indicate that Q. suber is generally well genetically equipped to handle climatic change in most of its current distribution (with the notable exception of Catalonia). Despite cork oak's long generation time, it seems reasonable that during the considered time frame current populations are able to shift their allele frequencies (2%– 12% on average, depending on the predictive model) due to a relatively high standing genetic variation, which according to (Kremer et al., 2012) should really work in the species’ favor in the presence of strong selective pressures.
This study, however, is limited to the considered environmental variables. Other factors that were not included in this work may have a larger effect on Q. suber's RONA. Inferring future adaptive potential of species is not yet commonplace practice (Jordan, Hoffmann, Dillon, & Prober, 2017; Rellstab et al., 2016); however, combining this type of study with ecological niche modeling approaches has the potential to greatly improve the accuracy of both kinds of predictions.
4.4 Final remarks
In this study, new nuclear markers were developed to shed new light on Q. suber's evolutionary history, which is important to understand, in order to attempt to predict the species response to future environmental pressures (Kremer et al., 2014).
Despite the relatively large geographic distances involved, the nuclear markers used in this work indicate a lesser genetic structuring than previously thought from cpDNA markers, which clearly segregated the species in several well-defined demes (Magri et al., 2007). The SNP data from this work can thus be used to propose two new hypotheses to replace the current view of a deep genetic structure as evidenced by cpDNA. The observed genetic structure can be explained either by balance between gene flow and local adaptation, or alternatively, differential hybridization of Q. suber with Q. ilex s.l. in the West and Q. cerris in the East being responsible for geographic differences’ origin, which are then maintained by the mentioned balance between gene flow and local adaptation (albeit more research is required to confirm this second hypothesis).
Despite the genetic structure homogeneity, outlier and association analyses hint at the existence of local adaptation. The RONA analyses suggest that this balance, between local adaptation and gene flow, may be a key in Q. suber's response to climatic change. It is also worth considering that despite the species’ likely capability to shift its allele frequencies for survival in the short term, the effects of such changes in the long term can be quite unpredictable (Feder, Egan, & Nosil, 2012; Lenormand, 2002), and only very recently have they began to be understood (Aguilée, Raoul, Rousset, & Ronce, 2016).
This study starts by providing a new perspective into the population genetics of Q. suber, and, based on this data, suggests an initial conjecture on the species’ future, despite the used technique's limitations. Even though studies regarding Q. suber's response to climatic change are not new (Correia et al., 2017; Vessella et al., 2017), this is the first work where this response is investigated from an adaptive perspective.
The software, pyRona, was developed and is provided in hopes that the method is adopted by the larger scientific community to estimate the RONA for other species, and eventually, make an impact in determining species conservation strategies. In this regard, the RONA results can be used in informing assisted migration projects (Aitken & Bemmels, 2016). In the specific case of the cork oak, European commercial stocks can be expected to benefit from the introduction of trees (and therefore alleles) adapted to more extreme temperature and precipitation conditions. As for which ones, should be further studied, but the genes that were functionally explored in this work should provide a good starting point.
In the near future, it is expected that improvements are made to the RONA method. In particular, using more sophisticated association testing (including the use of multivariate methods) and combining this approach with ecological niche modeling should yield much improved insights into species’ response to climatic change. These changes should be supported by expanding the use of the method to other species, which have both genetic and climatic data available.
ACKNOWLEDGEMENTS
We would like to thank R. Nunes, A. S. Rodrigues, C. Ribeiro, and I. Modesto, for their help during sample collection. We would further like to thank the two anonymous reviewers for the very through feedback they have provided. Field and laboratory work, and bioinformatics platform were supported by Fundação para a Ciência e Tecnologia (FCT)—Portugal [grant numbers SOBREIRO/0036/2009 (under the framework of the Cork Oak ESTs Consortium) and UID/BIA/00329/2013 (2015-2018)]. F. Pina-Martins was funded by FCT [grant number SFRH/BD/51411/2011 (under the PhD program “Biology and Ecology of Global Changes,” Univ. Aveiro & Univ. Lisbon, Portugal)].
DATA ACCESSIBILITY
Raw GBS data are available on NCBI's Sequence Read Archive (SRA) as “BioProject” PRJNA413625. A docker image containing the analysis process, software, and “assembled” data is available in https://hub.docker.com/r/stunts/q.suber_gbs_data_analyses/. The software pyRona is available in gitlab, and mirrored on github.