2 The Nuclear Envelope – Structure and Protein Interactions
Abstract
The proteins of the nuclear envelope have been characteriszd in detail in non-plant systems, revealing a complex series of interactions between the inner and outer nuclear envelope, with the proteins of the nuclear pore complex and connecting to the nucleoskeleton and cytoskeleton. Many of the proteins described in animal and fungal systems have not been identified in plants; however, recent advances have resulted in the description of several key plant nuclear envelope proteins including members of the Sad1/UNC84 (SUN) domain family and recently the first Klarsicht/Anc-1/Syne-1 Homology (KASH)-domain protein. This chapter describes the emerging networks of protein interactions at the plant nuclear envelope, together with discussion of their likely functions. In particular, the important role of plant SUN domain proteins as a key component of a linker of nucleoskeleton and cytoskeleton complex is described together with prospects for future study of plant nuclear envelope interaction networks.
2.1 Introduction
The nuclear envelope (NE) plays a fundamental role in eukaryotic cells. By separating the contents of the nucleus from the cytoplasm and by gate-keeping the traffic into and out of the nucleus, it enables the functioning of a complex and sophisticated genome. In consequence, it has been the subject of considerable interest and a substantial knowledge base has been created for animal and yeast cells. This chapter aims to review the current state of knowledge for plants, where recent discoveries are opening the field for rapid advance.
2.2 Organization and Structure of the Plant Nuclear Envelope
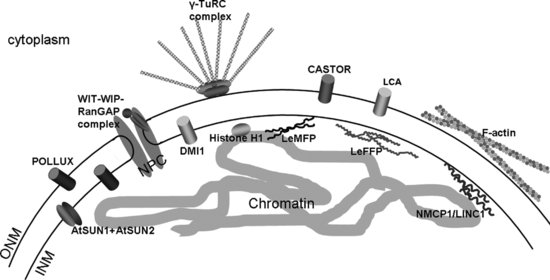
Plant nuclei are variable in shape and size and in vegetative tissue are often compressed into a narrow region of cytoplasm by a large central vacuole. However, in mitotic and undifferentiated cells, the nucleus is often rounded or ovoid, central and capable of movement within the cell cytoplasm. As in metazoans, chromatin is organized into regions of hetero- and euchromatin and there are great similarities in the overall organization of the internal structure of the nucleus with heterochromatin located adjacent to the NE. The shape and structure of the plant nucleus as well as its position are likely to be maintained by the interaction of the envelope with chromatin and the nucleoskeleton and with the cytoskeleton, just as in metazoans. The plant NE is also structurally similar to those of other kingdoms, with three identifiable domains: the outer nuclear membrane (ONM), the inner nuclear membrane (INM) and the pore membrane (Figure 2.1; Brandizzi et al., 2004). The ONM is linked to the perinuclear endoplasmic reticulum (PNER), functions in protein synthesis and is coated with ribosomes. The proteomes of the PNER and ONM are not identical and the junction region between the two membranes (diameter 2–30 nm) is believed to limit protein movement (Craig and Staehelin, 1988; Staehelin, 1997). Connections between the ONM and cytoskeleton are an important feature of the nucleus. Unlike higher metazoans, the entire plant ONM serves as a microtubule-organizing centre (MTOC) and plants lack centrosomes (Shimamura et al., 2004).
Biochemical Society Transactions 38, 307–311 © Portland Press Limited and International Federation for Cell Biology.
The second domain is the pore domain or pore membrane. As well as connecting INM and ONM and providing the environment in which the nuclear pore complexes (NPCs) are anchored, it helps to maintain the protein composition of the INM as membrane proteins entering the INM must pass through the periphery of the NPC, many requiring a nuclear targeting signal (King et al., 2006; Lusk et al., 2007). The structure and function of the NPCs and their relationship with the pore domain are dealt with in chapter The Plant Nuclear Pore Complex – The Nucleocytoplasmic Barrier and Beyond. The third domain, the INM, is closely associated with chromatin via numerous protein connections as well as with the ONM by protein bridges spanning the periplasm. In addition, a nucleoskeletal meshwork, termed the lamina in animals, underlies the INM, thereby supporting the structure and shape of the NE and the entire nucleus. To date, only two proteins have been characterized at the plant INM – Sad1/UNC84 (SUN) domain proteins. Hence the protein components that mediate most of the anchorage and functions at the plant INM remain to be identified (Figure 2.1).
Study of the plant NE reveals that while it is structurally similar, many of the key proteins providing these connections are absent. For instance, in metazoans, the lamina consists of filamentous lamin proteins. However, homologues of these are not present in the plant genome (Brandizzi et al., 2004). In addition, many other proteins interlinking the INM and chromatin are absent. These include homologues of animal and yeast NE proteins including members of the LEM1 (lamin–emerin–man1) domain family of proteins and the lamin B receptor (LBR). It may therefore be concluded that the plant NE is homologous to that of metazoans in structure and many of its functions; but is distinct in many of the protein interactions through which these features are achieved.
2.3 Proteins of the Plant Nuclear Envelope
The plant NE proteome has yet to be fully characterized, though recently, significant advances have been made in identifying plant NE proteins. This is partly due to the difficulty in isolating and purifying NE and partly due to the fact that such isolated NE is commonly associated with PNER; indeed as indicated above, the ONM and PNER proteomes at least overlap.
2.3.1 Proteins Involved in Signalling
One of the first proteins to be described at the plant NE was a calcium pumping ATPase of the sarcoplasmic reticulum/endoplasmic reticulum (SERCA) type called LCA (Figure 2.1; Downie et al., 1998). This first indication of a role for the NE in calcium signalling has since been corroborated by the identification of a NE Ca2+ signalling pathway involved in mycorrhizal infection and nodulation with the nuclear periplasm acting as a Ca2+ signalling pool (Chabaud et al., 2011). Components of this pathway described in the legume Medicago truncatula include the cation channel Doesn't make infection 1 (DMI1) in the nuclear membrane (Figure 2.1; Peiter et al., 2006; Riely et al., 2006); DMI2, a leucine-rich repeat (LRR) receptor-like kinase (Endre et al., 2002) and a calcium- and calmodulin-dependent kinase DMI3 in the nucleus (Mitra et al., 2004; Smit et al., 2005). Nodulation factors produced by the infecting organism trigger Ca2+ spikes in the nucleus (Oldroyd and Downie, 2006), detected by DMI3 and resulting in altered transcription required for successful symbiosis (Gleason et al., 2006). Two homologues of DMI1, Castor and Pollux, have also been reported in Lotus japonicus and also localize to the NE (Figure 2.1; Charpentiere et al., 2008). In addition, in silico studies predict the presence of various other Ca2+-sensitive and voltage-gated cation channels as well as ion pumps in both INM and ONM and suggest the plant NE plays an inherent role in signal transduction events to regulate gene activity (Matzke et al., 2009).
2.3.2 Proteins of the Nuclear Pore Complex
Further significant advances have recently been made in the identification of plant nucleoporins (Tamura et al., 2010). While structure and protein traffic at the NPC is considered in detail elsewhere in this volume, it is useful here to summarize the known components of the NPC. The work of Tamura and co-workers reveals that there are at least 29 nucleoporins present in Arabidopsis, of which 22 had not previously been described in plants (Tamura et al., 2010). Overall, the plant nups show a higher homology with vertebrate than yeast, commensurate with the greater size and complexity of function of the plant NPC (Nicotiana tabacum [tobacco] NPC ∼ 105 nm diameter, compared with vertebrate 110–120 nm and yeast ∼ 95 nm; Kiseleva et al., 2004; Fiserova et al., 2009). Orthologues of six vertebrate nucleoporins are not found in plants (Nup358, Nup188, Nup153, Nup45, Nup37, and Pom121). The major membrane anchor for the NPC found in vertebrates (but not yeast), gp210, is present in plants and is therefore a major component of the pore domain. Like its animal counterparts, gp210 has a single C-terminal transmembrane domain. While its molecular functions are yet to be studied, it has been identified in a screen of essential Arabidopsis genes to be indispensable for embryo development (Meinke et al., 2008). Two other plant nucleoporins (ALADIN, and Nup43) also have vertebrate, but not yeast, orthologues, which Tamura et al. (2010) suggest imply higher eukaryotic functions. Both are WD-repeat proteins likely to be involved in multiprotein complex formation.
Vertebrate NPCs have a cytoplasmic meshwork of filaments of importance in transport through the pores, which is driven by a RanGTP/ RanGDP gradient (Clarke and Zhang, 2008). Development of this gradient is defined by localization of two accessory proteins, Ran GTPase-activating protein (RanGAP) and the nucleotide exchange factor RCC1, which so far has not been identified in plants. In mammals, RanGAP is anchored to the cytoplasmic filaments by binding the small ubiquitin-related modifier protein (SUMO) followed by binding of this SUMOylated domain to Nup358. As the cytoplasmic filaments (Fiserova et al., 2009) and Nup358 are not present in plants (Xu and Meier, 2008; Tamura et al., 2010), the plant NE therefore has an alternative mechanism for localizing RanGAP. Attachment of plant RanGAP to the NE involves a plant-specific targeting domain, termed the WPP (tryptophan proline proline) -domain (Rose and Meier, 2001). This N-terminal domain is found in plant RanGAPs, plant NE-assocaited WPP proteins WPP1 and WPP2 as well as matrix attachment factor 1 (MAF1). The domain is necessary for RanGAP targeting and sufficient to target the heterologous protein GFP to the plant nuclear rim (Rose and Meier, 2001). Plant RanGAP is anchored to the NE by associating with WPP-interacting proteins (WIPs) and WPP interacting tail anchored proteins (WITs), which show predicted NPC localization (Figure 2.1; Xu et al., 2007; Zhao et al., 2008). Plant WIPs and WITs contain a C-terminal TM domain, which anchors the proteins to the NE membranes. A coiled-coil domain at the cytoplasmic side is thought to mediate both dimerization and association with the WPP domain of RanGAP. A triple knockout of WIPs 1–3 abolishes NE anchorage of RanGAP in undifferentiated root tip cells whereas RanGAP anchorage in other tissue seems to be independent of WIP and WIT making this a tissue- and development-specific anchorage process (Xu et al., 2007).
2.3.3 Proteins of the INM
The metazoan INM possesses an array of proteins involved in connecting the INM to the underlying nuclear lamina and to chromatin. Description of these proteins has been driven to a great extent by diseases such as progerias, skeletal defects, muscular dystrophies, lipoatrophy, and epilepsy resulting from INM protein mutations (Stewart et al., 2007). These proteins perform a variety of important tasks, some specific to vertebrates, but others found in all metazoans; their absence from the plant genome raises a number of important questions, given that INM proteins are implicated in nucleic acid metabolism, signal transduction, NPC spacing and the tethering of nuclear matrix and chromatin. Most of these functions also involve the lamina, which is tightly associated with the INM (Gruenbaum et al., 2005). Proteomic studies of the rat liver NE identified over 60 NE transmembrane proteins (Schirmer et al., 2003; Schirmer and Gerace, 2005).
The lamin B receptor (LBR) (Worman et al., 1988) is a key link between the nuclear lamina and INM. It has eight transmembrane domains, forms a multimeric protein complex, shows sterol reductase activity and contains a Tudor domain (involved in histone binding) (Huang et al., 2006). The N-terminus is located in the nucleoplasm and binds to lamin B and chromatin. Both LBR and its binding partners are absent from plants. In a recent review, Olins et al. (2010) provide evidence that LBR is conserved in organisms from the vertebrates including avians and amphibians to man, though it has not been detected in lower organisms and (of particular importance for this chapter) plants. Given the range of functions for LBR (reviewed by Olins et al., 2010) in post-mitotic NE reformation, interphase NE growth and heterochromatin positioning in interphase, we suggest that its roles must be undertaken by other plant proteins. The search for INM - chromatin bridges is therefore of high priority. Interestingly, while LBR is not present in plants, a construct derived from the N-terminus of the mammalian LBR including the first transmembrane domain and a nuclear localization signal (NLS) faithfully targets to the plant INM (Irons et al., 2003) and this has been used to investigate INM targeting in plants (Graumann et al., 2007; Graumann and Evans, 2011).
A further major group of proteins not present in plants is the LEM domain family. The LEM domain proteins are named after a conserved domain comprising lamina-associated polypeptide 1 (LAP1) and LAP2 in vertebrates; emerin, identified through its role in X-linked recessive Emery–Dreifuss muscular dystrophy and Man1 (Gruenbaum et al., 2005). The LEM-domain family tether chromatin to the NE (Gruenbaum et al., 2005). They bind directly to lamins and to barrier-to-autointegration factor (BAF), which binds to lamins and chromatin (Gruenbaum et al., 2005). Emerin also links nuclear actin with the INM and MAN1 acts as an antagonist in the transforming growth factor beta (TGFβ) signalling cascade (Gruenbaum et al., 2005; Bengtsson, 2007). Amino acid sensor independent 1-3 (ASI1-3, another LEM family member) are signal transducers that regulate expression of amino-acid permeases (Zargari et al., 2007).
The presence of a nucleoskeletal meshwork underneath the INM and chromatin associations at the NE suggests that the plant INM also contains proteins, which are involved in chromatin and nucleoskeletal anchorage and hence implied in controlling nuclear activities. It is therefore of great importance to identify these components and study their functional relationships with the nucleoskeleton and chromatin. To date two INM intrinsic proteins – the SUN domain proteins AtSUN1 and AtSUN2 have been identified and their functions are discussed in following sections.
2.3.4 Proteins Spanning the Periplasm and Linking the NE Membranes
The Linker of Nucleoskeleton and Cytoskeleton (LINC) complex (Crisp et al., 2006) is a multifunctional protein bridge that connects INM and ONM and provides anchorage for a variety of NE associated proteins in the nucleoplasm and cytoplasm supporting, moving and shaping the NE (Figure 2.2). In non-plant systems it is made up of two membrane integral components; SUN domain proteins (in the INM), ubiquitously present in higher and lower eukaryotes (Figure 2.3; Starr, 2009; Graumann et al., 2010) and KASH domain proteins of the ONM. The two families interact through their respective SUN and KASH domains in the periplasmic space separating the two membranes. While SUN-domain proteins show structural and sequence conservation across species (Figure 2.3), KASH-domain proteins are more diverse, permitting interactions with a wide range of cytoskeletal elements (Starr and Fridolfsson, 2010). In yeast, SUN domain proteins Sad1 (Schizosaccharomyces pombe) and Mps3 (Saccharomyces cerevisiae) are located at spindle pole bodies (SPBs) in addition to the NE (Bupp et al., 2007). Putative plant homologues of SpSad1 were first suggested in Arabidopsis (van Damme et al., 2004), and Oryza sativa (rice; Moriguchi et al., 2005) at the phragmoplast and mitotic spindle. The first detailed characterization by the authors (Graumann et al., 2010) revealed them to be classical SUN-domain proteins and showed that AtSUN1 and AtSUN2 are localized to the NE in interphase and provide first evidence of a putative LINC complex in plants (Figure 2.2). The SUN-domain proteins are considered in detail later in this chapter.
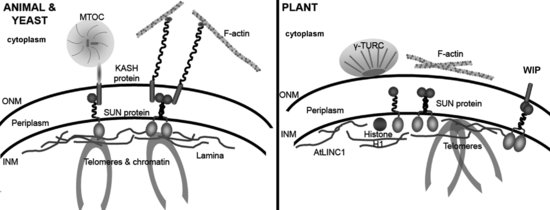

The KASH domain is a highly hydrophobic domain comprising a transmembrane region and a C-terminal KASH domain of 6-35 amino acids in the nuclear periplasm, which interacts with the SUN domain (Razafsky and Hodzic, 2009; Starr and Fridolfsson, 2010). Other than a short extreme C-terminal PPPX motif essential for SUN binding, the KASH domain does not show sequence homology sufficient for detection using a conventional bioinformatics approach and this short sequence is not found in plants. The cytoplasmic region of the KASH proteins is highly variable, though commonly containing spectrin repeats or coiled coils. Four KASH domain proteins have been characterized in mammals (nesprins 1–4) and in Caenorhabditis elegans (ANC-1, UNC-83, ZYG-12 and KDP), two in Drosophila melanogaster (Klarsicht and MSP-300) and in S. pombe (KMS1-2) and one in S. cerevisae (Cms4) (Wilhelmsen et al., 2006; Roux, et al., 2009; Starr, 2009). Each anchors specific cytoskeletal elements; nesprins 1 and 2 bind actin, nesprin 3 intermediate filaments and nesprin 4 kinesin (McGee et al., 2009; Starr, 2009). C. elegans, UNC-83 mediates the movement of the nucleus using kinesin (McGee et al., 2009; Meyerzon et al., 2009) whereas ZYG-12 interacts with dynein and is involved in microtubule organization and nuclear positioning in gonads (Zhou et al., 2009). Nesprins 1 and 2 are involved in nuclear migration during neurogenesis and neuronal migration in mice involving attachment of the nucleus to centrosomes by dynein/dynactin and kinesin (Zhang et al., 2009).
Very recently, the WIP protein family were identified to be structural and functional, but not sequence, homologues of KASH proteins in plants. As mentioned, they also consist of a cytoplasmic coiled-coil domain, are C-terminally anchored to the ONM and have a short, nine amino acid C-terminal tail. While this tail has no sequence homology to animal and yeast KASH tails, it contains a penultimate proline residue embedded in a VVPT motif that is conserved in other plant species (Zhou et al., 2012). In Arabidopsis, the WIP proteins have been shown to interact with SUN proteins and both the SUN domain and the VVPT motif are required for this interaction (Figure 2.2; Zhou et al., 2012). Evidence that WIP proteins are functional KASH homologues comes from observations that both WIP and SUN proteins are required to maintain nuclear shape. Knocking out either protein family results in rounded nuclei in various plant tissues (Zhou et al., 2012). In addition, the plant SUN-WIP complexes are involved in WPP-mediated RanGAP anchorage in undifferentiated root tip cells as knock out of AtSUN1 and AtSUN2 results in loss of RanGAP from the nuclear periphery (Zhou et al., 2012). This implies that LINC complexes can mediate plant-specific functions. Whether the WIP proteins are the only plant KASH proteins and whether they mediate NE-cytoskeleton interactions like their animal and yeast counterparts, remains to be elucidated.
2.3.5 The Plant Lamina
The presence of a meshwork of proteins, underlying and closely associated with the INM, is a feature of animal (Gruenbaum et al., 2005) and plant (Fiserova et al., 2009) nuclei. While the lamina of animal cells has been well characterized, that of plants is much less well described. The lamina of animal cells consists of lamins, type-5 intermediate filament proteins, together with lamin-associated proteins (reviewed in Wilson and Berk, 2010). Lamins are encoded by three genes, A, B1 and B2, which also generate the alternative splice product lamin C. The lamins are 60–80 kDa proteins and possess a characteristic structure with a long rod domain made up of linked α-helices and a highly conserved, globular N-terminus. A phosphorylation domain acting at the onset of NE breakdown controls association with lamin binding proteins. Plants lack sequence homologs of mammalian lamins (Brandizzi et al., 2004; Meier, 2007; Graumann and Evans, 2010a). However, there is now considerable evidence for a protein meshwork underlying the plant INM and attached to it (Minguez and Diaz de la Espina, 1993; Masuda et al., 1997; Fiserova et al., 2009). Recently, high-resolution field-emission scanning electron microscopy (Fiserova et al., 2009) has provided images revealing a complex filamentous structure at the nucleoplasmic face of the INM. Two types of filaments differing in thickness, 10–13 nm and 5–8 nm, were observed to interconnect to the NPC. The structure was also visible from the cytoplasmic face in detergent extracted nuclei and resembled the lamina meshwork of frog oocytes.
Several approaches have suggested the protein composition of the plant ‘lamina’. Initially, immunological studies revealed lamin-like nuclear intermediate filament-like proteins (McNulty and Saunders, 1992; Minguez and Diaz de la Espina, 1993; Masuda et al., 1997). The first of these proteins to be characterized was Daucus carrota (carrot) nuclear matrix constituent protein 1 (NMCP1), a 134 kDa NE-associated protein. Although much larger than mammalian lamins, NMCP1 has a central coiled-coil domain and head and tail domains, including a putative NLS (Masuda et al., 1997). There are NMCP proteins present in other dicots including Arabidopsis and Apium graveolens (celery) as well as monocots such as rice (Table 2.1; Moriguchi et al., 2005). As well as structurally resembling lamins, NMCP1 also has similar isoelectric point (5.6-5.8) and kinase-recognition domains (Masuda et al., 1997).
| Identity | ||||||
|---|---|---|---|---|---|---|
| Accession | Protein names | Organism | Length | (%) | Score | E value |
| D2YZU5 | Nuclear matrix constituent protein 1 | Apium graveolens | 1171 | 100.00 | 5884 | 0 |
| O04390 | Nuclear matrix constituent protein 1 | Daucus carota | 1119 | 84.00 | 4688 | 0 |
| A6BME3 | Nuclear matrix constituent protein 1-like | Petroselinum crispum | 1119 | 83.00 | 4673 | 0 |
| A6BME2 | Nuclear matrix constituent protein 1-like | Foeniculum vulgare | 1119 | 83.00% | 4665 | 0 |
| A6BME0 | Nuclear matrix constituent protein 1-like | Apium graveolens | 1119 | 83.00 | 4659 | 0 |
| A6BME1 | Nuclear matrix constituent protein 1-like | Coriandrum sativum | 1003 | 84.00 | 4190 | 0 |
| B9N1Z9 | Predicted protein | Populus trichocarpa | 1156 | 51.00 | 2698 | 0 |
| E5GCT1 | Nuclear matrix constituent-like protein 1 | Cucumis melo subsp. melo | 1205 | 48.00 | 2613 | 0 |
| A5BQE9 | Putative uncharacterized protein | Vitis vinifera | 1234 | 46.00 | 2450 | 0 |
| F4HRT5 | Protein little nuclei1 | Arabidopsis thaliana | 1132 | 43.00 | 2240 | 0 |
| B9SEG9 | ATP binding protein, putative | Ricinus communis | 1172 | 44.00% | 2219 | 0 |
| Q7XXP7 | Os02g0709900 protein (Putative nuclear matrix constituent protein 1) | Oryza sativa subsp. japonica | 1155 | 32.00 | 1574 | 1.00E–172 |
| D2YZU8 | Nuclear matrix constituent protein 2 | Daucus carota | 927 | 30.00 | 1046 | 1.00E–111 |
| D2YZU6 | Nuclear matrix constituent protein 2 | Apium graveolens | 925 | 30.00 | 1016 | 1.00E–107 |
| B9SX77 | Filamin-A-interacting protein, putative | Ricinus communis | 1052 | 30.00 | 992 | 1.00E–104 |
| Q0WQM6 | Putative nuclear matrix constituent protein | Arabidopsis thaliana | 743 | 33.00 | 963 | 1.00E–101 |
| F4JXK1 | Branched-chain-amino-acid aminotransferase 5 | Arabidopsis thaliana | 1010 | 29.00 | 952 | 1.00E–100 |
| Q9FLH0 | Putative nuclear matrix constituent protein 1-like protein (NMCP1-like) | Arabidopsis thaliana | 1042 | 29.00 | 940 | 1.00E–98 |
Using a reverse genetics approach, Dittmer et al., (2007) were able to characterize the Arabidopsis NMCP homologs they named ‘Little Nuclei’ or LINC. In addition to the structural characteristics suggested by sequence homology with NMCP1 and 2 (Table 2.1), T-DNA insertional double mutants of LINC1 and LINC2 showed significantly smaller nuclei (50% of wild type) as well as reduced plant growth. In addition, the mutant nuclei were more spherical. Expression of LINC1-GFP on its native promoter was high in meristematic tissues, but not differentiated tissues, such as mature root hairs and epidermal cells and the protein was localized to the periphery of the nucleoplasm (Dittmer et al., 2007). The LINC proteins therefore appear to be strong candidates as components of a plant nucleoskeleton; however, progress still needs to be made to prove that they constitute major components of the fibres observed in the electron micrographs or that they interact with proteins of the NE and with chromatin.
Other potential components of the higher plant nucleoskeleton have also been suggested, although without verification. A family of coiled-coil proteins, the filament-like proteins (FPP), was suggested as a possibility. The Lycopersicon esculentum (tomato) homolog, LeFPP, was identified in a yeast-two hybrid screen interacting with LeMAF1, an NE-associated protein involved in RanGAP binding the plant nucleus (Gindullis et al., 1999; Gindullis et al., 2002). In addition, a homolog from Pisum sativum (pea) was identified using a lamin B antibody (Blumenthal et al., 2004). There are seven family members of the FPPs in Arabidopsis (FPP1–FPP7; Gindullis et al., 2002) ranging from 615 to 1054 amino acids with four conserved motifs and a long coiled coil core. They remain to be functionally characterized.
Moriguchi et al., 2005, attempted to characterize the potential nucleoskeleton of rice. In addition to identifying a rice homologue of NMCP1 (see above), they also identified a number of other potential nucleoskeleton associated components. Ankyrin is a structural protein usually located at the plasma membrane; however, Nalp1, a 28 kDa rice ankyrin homologue, was located close to the INM, in a region termed the inner nuclear matrix region or chromatin-poor space. AtAnkyrin3, the Arabidopsis counterpart of Nalp1, shows high homology with the ankyrin repeat region of Nalp1 and is of similar size (27 kDa); they are however, much smaller than the human ankyrins and unlikely to perform similar functions (De Ruijter et al., 2000).
As well as proteins of the nucleoskeleton linking to the NE, there is also evidence for a variety of proteins involved in the interaction of the nucleoskeleton with chromatin domains. Plant nuclei contain actin, which forms nucleoplasmic microfilaments and associates with transcription sites (Cruz et al., 2008; Cruz and Moreno Diaz de la Espina, 2009). While the major actin binding proteins of the nucleoskeleton are absent (lamins, lamin associated proteins, nesprins), three actin-binding proteins have been identified (Perez-Muniva and Moreno Diaz de la Espina, 2011); profilin (Kandasamy et al., 2002), nuclear myosin 1 (Cruz et al., 2008; Moreno Diaz de la Espina, 2009) and actin depolymerizing factor (Ruzicka et al., 2007). Their molecular role in maintaining nuclear architecture remains to be determined.
The presence of spectrin-like proteins in pea nuclei was suggested by a study by De Ruijter et al. (2000) using antibodies raised to erythrocyte spectrin. Western blots revealed bands of 220–240 Da, similar to erythrocyte spectrin, and a prominent 60 kDa band. In light microscope immunolocalization, the antibodies detected proteins in puncta in the nucleoplasm and with staining increasing as the nuclear matrix was extracted progressively with detergents, DNase I and RNase A, and high salt. The authors suggest the data shows the presence of spectrin-like epitopes in plant nuclei, and postulated a role in stabilizing interchromatin domains. Recently, Pérez-Munive and Moreno Diaz de la Espina (2011), also using an immunocytochemical approach, have provided further evidence for spectrin chain proteins that co-immunoprecipitate and co-localize with nuclear actin after detergent extraction of the nuclear matrix of onion (Alium cepa) with nuclease digestion. Using antibodies to chicken spectrin, they observed that spectrin-like proteins and plant intermediate filaments co-localize in the nucleoplasm suggesting a structural role. Spectrin-like proteins were shown to be distributed at or near the NE, associated with chromatin and at the nucleolus. The authors are investigating interactions between the spectrin-like proteins and NMCP family members; however, to date sequence analysis of the proteins do not indicate homology with animal spectrins (Pérez-Munive and Moreno Diaz de la Espina, 2011).
An alternative approach to the identification of putative plant lamins resulted from the isolation of nuclear matrix proteins and use of an anti-intermediate filament monoclonal antibody MAb TIB 131 by several laboratories. In plants, researchers have shown that this antibody and various other anti-lamin antibodies recognize nuclear 60 and 65 kDa proteins (McNulty and Saunders 1992; Minguez and Moreno Diaz de la Espina, 1993). In the most detailed of these studies (Blumenthal et al., 2004) proteins of 54, 60 and 65 kDa were suggested as putative nuclear intermediate filament proteins in pea. Limited sequence data from the 65 and 60 kDa proteins showed homology with intermediate filament proteins and antibodies raised to the 65 kDa protein also cross-reacted with the smaller proteins and with an avian lamin preparation. Immunocytochemistry suggested nucleoplasmic, but not specifically nuclear peripheral, localization. Negatively stained extracts of the fractions show filaments of 6–12 nm diameter.
2.4 The Plant Nuclear Envelope and the Nucleoskeleton; Attachments at the INM
As many of the key proteins involved in attaching the nucleoskeleton (like members of the LEM domain family) are absent from plants, attention has focussed on plant members of the SUN-domain family as key INM proteins with the potential to act as bridges between envelope and nucleoskeleton.
The SUN-domain proteins occur throughout the plant kingdom. Classical (C-terminal) SUN-domain proteins have been identified in the club moss, Selaginella meollendorffii, the moss, Physcomitrella patens and in algae (Table 2.2) as well as in a range of both monocot and dicot species. They have been characterized in detail in two species: Arabidopsis (Graumann et al., 2010; Graumann and Evans, 2011; Oda and Fukuda, 2011) and Zea mays (maize; Murphy et al., 2010). In each case, two proteins with a C-terminal SUN domain have been described, designated AtSUN1 and AtSUN2 (Graumann et al., 2010) and ZmSUN1 and ZmSUN2 (Murphy et al., 2010). Sequence comparison of both entire sequences (Table 2.2) and of the SUN domains (Figure 2.3) reveal that they show a greater homology within the monocots and within the dicots than between the two groups; for instance AtSUN1 and AtSUN2 show a higher degree of homology (68% identity, 1.00 E−178) with each other than with either ZmSUN1 or ZmSUN2 (41%, 4.0 E−79 and 2.0 E−70 respectively). AtSUN1 is located on chromosome 5, AtSUN2 on chromosome 3, while in maize, ZmSUN1 is located on chromosome 5, and ZmSUN2 is located on chromosome 3.
| SUN Domain | Annotated | |||
|---|---|---|---|---|
| Uniprot | Species | (pfam) | Name | Length |
| Proteins with a predicted C-terminal SUN domains | ||||
| Q9FF75_ARATH | Arabidopsis thaliana | 313-415 | AtSUN1 | 471 aa |
| Q8L9I5_ARATH | Arabidopsis thaliana | 140-276 | 285 aa | |
| Q9SG79_ARATH | Arabidopsis thaliana | 310-446 | AtSUN2 | 455 aa |
| A9TD16_PHYPA | Physcomitrella patens | 145-288 | 293 aa | |
| A9RVG5_PHYPA | Physcomitrella patens | 152-295 | 300 aa | |
| A2WN90_ORYSI | Oryza sativa indica | 303-444 | 455 aa | |
| Q5NBL8_ORYSJ | Oryza sativa japonica | 303-444 | 455 aa | |
| B6TRH2_MAIZE/ B6TY16_MAIZE | Zea mays | 289-427 | ZmSUN2 | 439 aa |
| C5XGK7_SORBI | Sorghum bicolour | 15-115 | 126 aa | |
| A2Y2L0_ORYSI | Oryza sativa indica | 309-449 | 453 aa | |
| Q7XXP5_ORYSJ | Oryza sativa japonica | 309-449 | 453 aa | |
| C5XW88_SORBI | Sorghum bicolour | 295-435 | 443 aa | |
| B6THU8_MAIZE | Zea mays | 314-454 | ZmSUN1 | 462 aa |
| B9SBM5_RICCO | Ricinus communis | 322-460 | 471 aa | |
| A5BJ94_VITVI | Vitis vinifera | 320-364; 532-629 | 640 aa | |
| B9HVF3_POPTR/ B9HK83_POPTR | Populus trichocarpa | 148-286 | 292 aa | |
| A4RXN9_OSTLU | Ostreococcus lucimarinus | 1534-1671 | 1676 aa | |
| C1MM01_MICPC | Micromonas pusilla | 830-967 | 683 aa | |
| C1E249_MICPC | Micromonas pusilla | 381-516 | 526 aa |
Plant C-terminal SUN-domain proteins are smaller than their metazoan counterparts (Figure 2.3), the plant proteins having between 430 and 480 residues (yeast SAD1 with 524 residues). There is a strong structural resemblance between the plant C-terminal SUN domain proteins (Figure 2.3) and with the mammalian and yeast proteins, with a single transmembrane domain located towards the N-terminus and a coiled-coil domain between the transmembrane domain and SUN domain. In common with the non-plant family members, the N-terminus is located in the nucleoplasm and the coiled-coil domain and SUN domain in the nuclear periplasm (Graumann et al., 2010; Graumann and Evans, 2010b).
Evidence that the plant SUN domain proteins are involved, directly or indirectly, in connections between the envelope and nucleo- or cytoskeleton comes from a number of experimental approaches (Figure 2.2). A number of groups have generated double T-DNA mutant plants for AtSUN1 and AtSUN2 (Armstrong and Osman, unpublished; Zhou et al., 2012); Oda and Fukuda (2011) incorporated a micro (mi) RNA interference construct to improve knockdown efficiency. While they observed no significant major phenotypic differences, for instance in plant growth, development or fertility, they did observe that typical elongated nuclei of root epidermis and root hairs of wild-type plants were rounded in the SUN knockdowns. Movement of the nucleus in developing root hairs was, however, not affected. As interactions between INM proteins and nucleoskeletal elements such as the lamins are known to regulate nuclei shape, it suggests that the roles of AtSUN1 and AtSUN2 in maintaining nuclear shape may also involve interactions with plant nucleoskeletal elements. Further evidence for this comes from a study in which knock down of the lamin-like LINC proteins resulted in similarly spherical nuclei (Dittmer et al., 2007).
Studies of the behaviour of SUN domain proteins in mitosis also strongly points to their association with the nucleoskeleton and cytoskeleton. The NE of higher plants breaks down in what is termed ‘open mitosis’ and reforms around decondensing chromatin at the end of division. This process involves the breaking of protein bridges between the membrane and its associated proteins, followed by reformation of these links. The SUN-domain proteins in metazoans are among the first proteins to reassociate with chromatin. In tobacco suspension culture cells, AtSUN1 and AtSUN2 migrate to mitotic ER membranes at envelope breakdown and then rapidly accumulate in the reforming NE at the end of division, strongly suggesting direct or indirect association with chromatin. Plant NE reformation is spatially organized as AtSUN1 and AtSUN2 first aggregate at the surface of chromatin facing the spindle pole, then accumulate at the sides and finally facing the cell plate (Figure 2.4; Graumann and Evans, 2011). It has been shown that the lamin-like NE associated NMCP1 and NMCP2 in celery cells behave similarly (Kimura et al., 2010). It is interesting that AtSUN2 accumulates at the NE specifically during prophase and a role in chromatin organization prior to NE breakdown has been suggested (Graumann and Evans, 2011). In a parallel study, Oda and Fukuda (2011) explored the location and involvement of microtubules using a fluorescent tubulin construct with AtSUN1-mRFP. They observed microtubules both in association with the NE during NE breakdown and associated with the mitotic membranes. The SUN-domain proteins also localize to the phragmoplast – an area of high cytoskeleton and membrane activity forming the new cell wall and AtSUN2 may be interacting with other proteins here (Graumann and Evans, 2011; Oda and Fukuda, 2011).
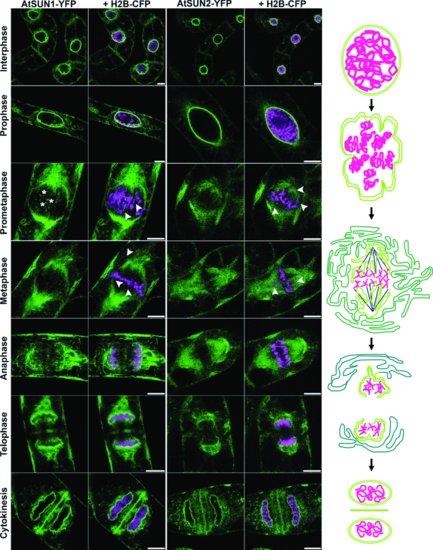
In addition to the classical SUN-domain proteins, recent in silico evidence and studies in maize suggest the presence of a second group of SUN-domain proteins, termed the mid-SUN proteins, where the highly conserved SUN domain is localized at the centre of the protein (Figure 2.3). Three of these mid-SUN proteins have been identified in maize, Arabidopsis and Chlamydomonas reinhardtii (Murphy et al., 2010; Graumann, Brickley and Tatout, unpublished observations). Immunological evidence suggests that the mid-SUN domain proteins are also localized at the NE. However, their functions and characteristics remain to be studied.
2.5 The Plant Nuclear Envelope and the Cytoskeleton; Attachments at the ONM
Initial evidence for the attachment of the NE to the cytoskeleton is indirect; in a number of circumstances, in particular in development and in response to stress, plant nuclei move in a manner dependent on the actin cytoskeleton (Chytilova et al., 2000; Ketelaar et al., 2002). Nuclear movement in response to abiotic and biotic stimuli such as bacterial, fungal and viral infections is considered later on in detail. Whether the connection between actin filaments and the NE is direct or mediated through protein complexes is unknown (Figure 2.2). Treatment of cells with latrunculin B or other actin depolymerizing drugs reduces nuclear movement in leaf epidermal cells expressing an NE marker (Graumann et al., 2007). While the full extent of nuclear movement has yet to be described, observations of nuclei in root epidermal cells and root hairs have been made by a number of groups (Chytilova et al., 2000; Ketelaar et al., 2002). Long-distance nuclear migration requires large actin bundles (Iwabuchi et al., 2010) while finer actin filaments position nuclei within fully elongated root hairs (Iwabuchi et al., 2010). The actin-associated motor protein myosin XII has also been found to localize to the plant NE (Avisar et al., 2009), indicating that not only actin bundles per se but also actin-associated proteins and motorproteins may play a role in nuclear movement. The presence of proteins for nucleating tubulin (Seltzer et al., 2007) at the ONM (Figure 2.2) suggests that the microtubule cytoskeleton might additionally play a role in nuclear size or positioning and it has been shown that destabilizing of microtubules increases the rate of movement of the nucleus (Fournier et al., 2008).
Less is known about the protein components by which the cytoskeleton is anchored to the NE. In non-plant systems, the γ-tubulin ring complex (γ-TuRC) forms the central part of microtubule organizing centres (MTOC) and SPB. The γ-TuRC comprises five γ-tubulin complex proteins (GCP) and γ-tubulin itself (Fava et al., 1999; Murphy et al., 2001). Two plant proteins, AtGCP2 and AtGCP3, have been shown to be homologs of Drosophila and yeast GCP2 and 3, and form a soluble complex with γ-tubulin that associates with the plant ONM (Seltzer et al., 2007). Both have NE targeting domains that are thought to target the γ-TuRC to the NE, which is then retained there by associating with an as yet unknown ONM intrinsic protein (Figures 2.1 and 2.2; Seltzer et al., 2007). In metazoan and yeast, MTOCs and SPB are anchored to the NE by KASH proteins. Similarly, actin filaments are associated with the NE by interacting with the actin-binding domain of KASH proteins (Figure 2.2). While WIP proteins have been identified as plant KASH homologs, it is unclear whether they participate in anchoring cytoskeletal elements to the NE. A triple knock-out mutant, which lacks all three WIP proteins results in rounded nuclei indicating these proteins are needed to shape nuclei (Zhou et al., 2012). Whether this process is also dependent on cytoskeletal elements or whether other plant KASH-like proteins mediate this remains to be elucidated.
2.6 Targeting of Proteins to the Plant NE
The molecular mechanisms of trafficking membrane-intrinsic proteins to the INM have predominantly been studied in metazoan and yeast systems while evidence of these processes in plants remains sparse. The endoplasmic reticulum (ER), ONM, pore membrane and INM are continuous, and therefore all proteins destined to reside in the INM pass through the pore membrane. This was thought to be due to passive diffusion, with the INM proteins retained by binding to nucleoplasmic components (Figure 2.5a; Mattaj, 2004). However, while proteins smaller than 25 kDa may enter the INM by this means, the process is slow and inefficient. It is now clear that INM-intrinsic proteins enter the nucleus using an energy-dependent mechanism (Ohba et al., 2004) similar to that for soluble proteins (see Chapter The Plant Nuclear Pore Complex – The Nucleocytoplasmic Barrier and Beyond), involving the Ran cycle (King et al., 2006; Lusk et al., 2007) and requiring an NLS. The INM proteins are first sorted at the translocon where association with importin-α occurs (Figure 2.5b). They are differentiated from other membrane proteins by their transmembrane domains (Saksena et al., 2004; Saksena et al., 2006; Braunagel et al., 2007). Many INM proteins have an NLS, which is recognized by the import machinery and is partly essential for correct targeting to the INM (King et al., 2006; Lusk et al., 2007). Entry through the pore (probably the region of the pore membrane) requires changes in NPC structure, provided by flexible, sliding nucleoporins (King et al., 2006; Alber et al., 2007; Lusk et al., 2007). In addition, FG nucleoporins are required for INM protein translocation and some, such as POM121, are associated with, or are in close proximity to, the pore membrane (Alber et al., 2007; Lusk et al., 2007).
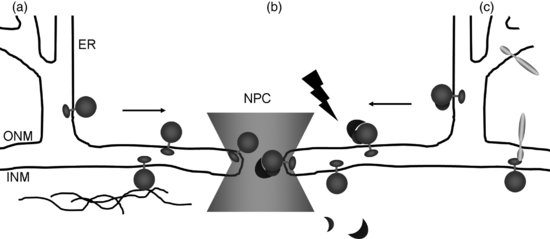
Targeting of NE proteins in plants is far less well understood. However, plants show similar mechanisms for the import and export of soluble proteins through NPC (Merkle, 2004; Meier, 2007; Meier and Brkljacic, 2009) with 17 importin-β, eight importin-α and three exportin homologs identified (Brandizzi et al., 2004; Meier, 2007). Most members of the Ran cycle have been characterized (Meier and Brkljacic, 2009) and putative nucleoporins with FG repeats have been identified (Tamura et al., 2010). The presence of a classical NLS has also been identiftied in plant NE proteins such as DMI1, AtSUN1 and AtSUN2 and the WIPs (Xu et al., 2007; Charpentier et al., 2008; Graumann et al., 2010). The fact that the N-terminus and first transmembrane domain of human LBR fused to GFP (green fluorescent protein; LBR–GFP) localizes to the INM in tobacco cells, also strongly suggests that mammalian INM targeting signals are recognized by plants (Irons et al., 2003; Graumann et al., 2007). Apart from the seemingly conserved INM targeting mechanisms, plants also have a specific NE targeting mechanism for NE-associated proteins (Meier, 2007). This system is based on the WPP domain, which is present in LeMFP1, MAF1 and its Arabidopsis homologues WPP1 and WPP2 as well as Arabidopsis RanGAP (Meier, 2007; Zhao et al., 2008). Whereas in animals RanGAP is anchored to the ONM by SUMOylation (SUMO is small ubiquitin-related modifier) and binding to RanBP2 (Ran-binding protein 2)/Nup385, in plants the WPP domain is essential for association with the ONM and NPC by interacting with WIPs and WITs (see Section 2.3.2.) (Meier, 2007). This RanGAP targeting event is specific to undifferentiated Arabidopsis root-tip cells, suggesting that this process is linked to development and is more complex than in animal cells (Meier, 2007).
Targeting of membrane proteins to the ONM appears more straightforward and does not require transfer through the pores. As the ONM and ER form a continuum, many proteins are present in both membranes. However, KASH proteins have been found to be specifically localized to the ONM. They are retained there by binding interactions with SUN proteins (Figure 2.5c). Thus, when the plant KASH homolog WIP1 is expressed in cells lacking AtSUN1 and AtSUN2, it no longer accumulates at the nuclear periphery (Zhou et al., 2012). Whether this binding retention mechanism is used by other ONM proteins is currently not known.
2.7 Nuclear Envelope Protein Dynamics in Mitosis
2.7.1 The Role of NPC in Regulating NE Dynamics in Cell Division
The first major NE event in dividing higher plants and metazoans occurs prior to NE breakdown (NEBD) in G2, when NPCs replicate, the NE enlarges and DNA is duplicated. During this, new NPCs are inserted, either by joining INM and ONM to create a pore into which the NPC proteins insert (requiring protein interaction with the membrane) or nucleoporins are inserted into an intact membrane and cause fusion of the ONM and INM, after which recruitment of the other nucleoporins follows (Hetzer, 2010). Membrane fusion occurs at the luminal face of the membranes to form the pore. Proteins are then added from both the nucleoplasmic and cytoplasmic face (D'Angelo et al., 2006). Drin et al. (2007) have shown that Nup133 contains an α- helical domain that senses membrane curvature. If sensing curvature by this domain is a prerequisite for development of the pore, it implies that membrane fusion and pore formation precede the construction of the NPCs (Hetzer, 2010). It is interesting to note that at this stage, as insertion of NPCs has preceded NEBD, the processes parallel those in eukaryotes with closed-cell division like yeast, where pore insertion has been characterized in detail. Here, the pore-domain transmembrane proteins (Pom34, Pom152 and Ndc1) mark the point of insertion of the NPC, possibly by bringing ONM and INM together. Nuclear pore assembly involves Nups59/53 with integral membrane proteins Pom34 and Pom152, to which Nup170 and membrane-integral nucleoporin Ndc1 attach (Onischenko et al., 2009). Depletion of Nup170 and Nup157 results in pores that are mis-located to the INM and cytoplasm rather than creating true pores (Dawson et al., 2009).
The insertion of NPCs into the plant NE in interphase has received little attention. In a study by Fiserova et al. (2009), NPCs in three–day old and ten-day old tobacco BY-2 suspension cultures were observed. The NPCs were highly abundant in all cells (40–50 per μm2). The authors suggest that the three-day old cells were likely to be in S phase. At this stage, NPCs were distributed over the nuclear surface, with around 30% of pores in connected pairs. In older cells, rows of NPC were observed instead. In addition, pores in three day old cells appeared simpler than those in older cells, and the cytoplasmic ring thinner, with structures similar to those observed in Xenopus leavis oocytes. However, to date, antibody and molecular probes have not been available to permit detailed sequential analysis of NPC formation in plants and the hypothesis that the processes involved are the same as those in animals or yeast has yet to be tested.
While plants possess the Ran cycle for transport at the pores (see above), the Ran nucleotide exchange factor ‘regulator of chromatin condensation 1’ (RCC1) is not present and RanGAP is attached to the NPCs by interaction via a C-terminal WPP domain with the WIPs and WITs (Rose and Meier, 2001; Jeong et al., 2005; Xu et al., 2007; Zhao et al., 2008). In addition to this role, Ran is also directly involved in pore assembly. In yeast, high levels of Ran-GTP and decreased importin stimulate pore formation (Harel et al., 2003; Walther et al., 2003). In xenopus oocytes, NE with nuclear pores assemble around beads coated with Ran in the presence of RCC1, forming pseudo-nuclei (Zhang and Clarke, 2000). Importin has also been shown to be essential to the process by which pores assemble on chromatin (Rotem et al., 2009). A role for Ran in NPC insertion in plants has yet to be considered.
Open mitosis requires the detachment of the envelope from its underlying structures (chromatin and the nucleoskeleton) and its breakdown and separation from chromatin. In animals, the NE membranes and some of its protein constituents migrate into the ER network, to form the ‘mitotic ER’ or ‘mitotic membranes’. Before these events, the NPCs are removed, giving exchange between nucleoplasm and cytoplasm. Pore removal is regulated and soluble NPC components migrate to the cytoplasm or become part of the mitotic apparatus, some remaining as protein complexes (Hetzer, 2010).
The first stage in the breakdown of NPCs is the removal of Nup98 (Griffis et al., 2002), followed by other nucleoporins (Dultz et al., 2008). In animal cells, this occurs in prometaphase while in plants it is earlier, in late prophase (Rose, 2007). Linkage of NPC to the lamina is implied for animals as mutation, down-regulation of expression, or introduction of Fab fragments of antibodies to gp210 in C. elegans results in failure of the lamin nucleoskeleton to depolymerize. Phosphorylation of the C-terminus of gp210 by cyclin B is essential for NEBD (Galy et al., 2008).
Following NEBD in plants, Tpr, which is associated with the nuclear pore basket, migrates to the mitotic spindle in prometaphase (Xu et al., 2007). Tobacco Rae1 associates with mitotic microtubules including the pre-prophase band (PPB), spindle and phragmoplast (Lee et al., 2009). Deletion or down regulation of the gene for Rae1 causes defects in the spindle organization, chromatin alignment and segregation as well as decreased levels of cyclin B, cyclin-dependent kinase B1-1 (CDKB1-1) and histones H3 (Lee et al., 2009).
Removal of NPCs has a number of consequences, including the loss of selective permeability of the membrane. Exposure of the nucleoplasm to cytoplasm initiates a series of phosphorylation events. Cyclin dependent kinases (CDKs) and cyclin B1 enter the nucleus and cause dissociation of proteins of the lamina and INM, which are then released into the ER. Breakdown then progresses as NE proteins move to the mitotic ER. Dephosphorylation at the end of division reverses this (Anderson and Hetzer, 2008; Guttinger et al., 2009). Progress of NEBD is regulated by aurora kinases and CDKs. How these kinases regulate plant NEBD and reformation remains to be established; however, aurora kinases are associated with the plant NE in interphase and localize to the mitotic spindle, centromeres and phragmoplast in division. They phosphorylate histone H3 and are involved in chromosome segregation and cytokinesis (Demidov et al., 2005; Kawabe et al., 2005). Plant B1 cyclins also accumulate at the NE (Rose et al., 2004). The down regulation of cyclin B and CDKB1-1 in combination with decreased mitotic activity in NbRae1 mutants suggests their involvement in plant mitosis progression (Lee et al., 2009). In addition to kinases, RanGAP plays a critical role in NEBD (Meier and Brkljacic, 2009).
Loss of physical interactions within the nucleus permits physical forces to tear and open the membrane. Microtubule-dynein interactions draw membrane away from the lamina (Beaudouin et al., 2002; Salina et al., 2002; Muhlhausser and Kutay, 2007; Stewart et al., 2007) although Lenart et al. (2003) suggest that NEBD can occur in the absence of a microtubule motor. The process in plants also involves positional information in order to establish the plane of division required in a walled structure, where the formation of a PPB of microtubules predicts the division plane. Tearing of the NE occurs where ONM and PPB are closest to one another and before the PPB disintegrates (Dixit and Cyr, 2002; Brandizzi et al., 2004; Rose et al., 2004; Evans et al., 2009).
2.7.2 NE Protein Dynamics in Division
One of the first studies of the location of a plant NE component in mitosis was in 1998, when Downie et al. (1998) used immunostaining to detect NE-specific staining of the Ca2+ ATPase LCA in spindle associated membranes of dividing tomato cells. Recently, the behaviour of plant NE components in mitosis has been observed using the chimaeric LBR-GFP probe described earlier (Irons et al., 2003), the plant SUN-domain proteins (Graumann and Evans, 2011; Oda and Fukuda, 2011) and the putative plant lamin-like proteins NMCP1 and NMCP2 (Kimura et al., 2010). As expected, both plant NE membrane markers and the putative plant lamin-like proteins disassemble at the beginning of mitosis (Figure 2.4). As the nucleoskeleton is lost in celery cells, NMCP1 associates with the mitotic spindle while NMCP2 migrates to the mitotic cytoplasm and both join the reforming NE at the end of the process (Kimura et al., 2010). While this study suggested that in fixed celery cells, dye-stained membranes from the NE formed vesicles, studies using live-cell imaging reveal a different behaviour, similar to that observed in metazoans. Human-derived LBR-GFP, in use as a plant NE marker, suggest that NE membranes in tobacco BY-2 cells were distributed to mitotic membranes (Irons et al., 2003; Graumann and Evans, 2011). This was also corroborated by studies using native plant proteins described below.
The native plant INM markers AtSUN1 and AtSUN2 also reveal that membranes of the NE enter the pool of mitotic membranes after NEBD (Graumann and Evans 2011; Oda and Fukuda, 2011). The SUN-domain proteins (like LBR-GFP) localize to spindle membranes and to tubules traversing the division zone in metaphase, while most of the spindle membranes accumulate at the spindle poles (Figure 2.4). In contrast with metazoans, where NE proteins distribute throughout the mitotic ER leaving metaphase chromosomes devoid of membranes, membranes containing NE proteins remain in close proximity to chromatin throughout division in plants (Anderson and Hetzer, 2008; Hetzer, 2010; Graumann and Evans, 2011). In metazoans, NE reformation commences with mitotic ER tubules containing INM proteins contacting decondensing chromatin in anaphase (Webster et al., 2009; Hetzer, 2010) with peripheral and core chromatin areas attracting different INM proteins (Güttinger et al., 2009). Both Graumann and Evans (2011) and Oda and Fukuda (2011) observed that NE reformation is spatially organized as AtSUN1-YFP and AtSUN2-YFP aggregate initially on chromatin facing the spindle pole followed by accumulation around the sides and finally localization of the proteins on chromatin facing the cell plate (Figure 2.4). This has not been observed in metazoans (Ellenberg et al., 1997; Webster et al., 2009; Hetzer, 2010) and may be plant specific, commensurate with a linkage to specific chromatin domains, and suggests a tight spatial regulation of NE reformation. Similar observations were made of NMCP1 and NMCP2. NMCP1 also first assembled on chromatin facing the spindle in late anaphase and at telophase completely surrounded the decondensing chromatin (Kimura et al., 2010). Thus AtSUN1 and AtSUN2 may be recruited to the reforming NE by interactions with chromatin or other specifically targeted nuclear components such as NMCP1.
2.8 The Phragmoplast and Cell Plate and their Relationship to the NE
The final process in the production of two daughter cells, accompanying the completion of the formation of the NEs, is the formation of the new dividing cell wall. This is generated along the plane of division predicted by the PPB microtubules and involves considerable secretory vesicle activity. Use of the mammalian-based LBR-GFP expressed in plants shows that it is not only present in the newly forming NEs, but also in the phragmoplast (Irons et al., 2003; Brandizzi et al., 2004; Evans et al., 2011). Other known NE proteins, including the WIPs and WITs (other than WIP3; Zhao et al., 2008) and the SUN-domain proteins (Graumann and Evans, 2011) are also observed in the phragmoplast (Figure 2.4). This may be due to a lack of an effective sorting and retrieval mechanism and to the very high volume of membrane traffic directed to the cell plate. Alternatively, it may suggest a role for these proteins in cell wall formation. Interestingly, RanGAP1 is specifically localized to the position of the PPB and remains in location throughout division during mitosis and cytokinesis (Xu et al., 2007). This suggests that a Ran gradient is formed that directs the vesicle traffic to the phragmoplast. As Ran has also been shown to be involved in the formation of the NE and insertion of NPCs, it seems possible that similar mechanisms are involved in both systems resulting in mis-direction of vesicle traffic. Alternatively, some NE proteins may be caught up in the rapid flow of membrane to the phragmoplast and are subsequently removed by recovery mechanisms. AtSUN2–YFP appears to have a significant immobile fraction at the cell plate indicating interactions and therefore may be functional (Graumann and Evans, 2011). It is interesting to observe that so far only in plants has spatial NE reformation from proximal to the spindle to proximal to the cell plate been observed, and it is intriguing to speculate whether this is associated with the presence of NE proteins in the cell plate. Among other factors such as kinases, phosphatases, Ran and chromatin remodelling components, the amount of available membrane is known to affect NE reformation (Webster et al., 2009) and it is possible that the cell plate and phragmoplast form membrane reservoirs that affect the distribution of membranes in the plant cell. Thus the SUN-domain proteins might be involved in such regulation.
2.9 The Plant NE in Meiosis
In addition to the events described for mitosis, the NE has a specific function in prophase 1 of meiosis as it anchors and positions telomeres. Anchorage of the telomeres to the NE is mediated by the SUN-domain components of the LINC complex in animal and yeast systems and deletion of these can result in abolition of gametogenesis (Tomita and Cooper, 2006; Ding et al., 2007). In animal, yeast and some plants a chromosome bouquet is formed and anchored to the NE via the telomeres. In animals and yeast this is essential for homologous pairing (Tomita and Cooper, 2006; Roberts et al., 2009). In some plant species, however, this structure is not well defined. For instance, in the model plant Arabidopsis, the telomeres are first clustered around the nucleolus and then move to the nuclear periphery (Roberts et al., 2009). How telomeres are linked to the nuclear periphery in plants remains to be investigated but the presence of plant SUN-domain proteins suggests that similar mechanisms may exist (Graumann and Evans, 2010b).
2.10 Lipid Composition of the Plant NE and its Homeostasis
Eukaryotic endomembrane bilayers consist mainly of phospholipids and glycolipids but also contain smaller amounts of sterols and sphingolipids (Jouhet et al., 2006; Aubert et al., 2011). Instead of being randomly located throughout the membrane, lipid composition of membranes is controlled by protein-protein and protein-lipid interactions and results in specific membrane domains (Jouhet et al., 2006; Aubert et al., 2011). Thus membrane lipids, in addition to membrane imbedded proteins, play a significant role in establishing and maintaining the structure and function of specific membranes. Consequently, the lipid composition of the plant NE is also non-random and highly controlled. Biochemical analysis of onion root and stem tissue found that the NE membranes consist of approximately 60% lecithin and between 20–24% phosphatidyl ethanolamine. Non-polar lipids such as sterols and triglycerides make up 35–45% of the NE lipid content. Indeed, the onion NE has a relatively high sterol content with sitosterol making up 80% of the total sterols. Finally, the study also found that 80% of the fatty acids in the membrane phospholipids are unsaturated, contributing to the fluidity of the membrane (Philipp et al., 1976).
Most of phospholipids, sterols and sphingolipids in endomembranes are synthesized in the ER and from there transported to other internal membranes or the plasma membrane. The physical continuity between ER and NE suggests that they may share a similar lipid composition. In support of this, Philipp et al. (1976) found that the rough ER (rER) and NE of onion cells have an identical phospholipid composition. It is therefore reasonable to think that the bulk of NE lipids are transported by lateral diffusion from the ER, through to the ONM, pore membrane and INM (Jouhet et al., 2006). Two other transport modes exist to disperse lipids from the ER – diffusion across the bilayer mediated by flipases and movement of individual lipids outside the bilayer through an aqueous phase (Jouhet et al., 2006). The latter method is mediated through membrane contact sites and lipid transfer proteins. Evidence from yeast and mammalian systems suggests such a transport mode is present at the NE and involved in NE lipid homeostasis (Jouhet et al., 2006; Mijaljica et al., 2010).
2.10.1 Nuclear-Vacuolar Junctions and Lipid Homeostasis
Membrane contact sites are areas where two membranes are in very close proximity – around 10 nm. These sites contain lipid transfer proteins, which can shuttle individual lipids through the aqueous phase between the two membranes (Jouhet et al., 2006). The membrane contact site at the NE consists of the ONM and the tonoplast of the lytic vacuole; hence they are referred to as nucleus-vacuole junctions (Kvam and Goldfarb, 2004; Jouhet et al., 2006). These junctions have been best studied in S. cerevisiae but also exist in mammalian cells. Whether they are also present in plants remains to be studied. However, one of the protein components of the nucleus-vacuole junctions has a homolog in Arabidopsis. The two main proteins that maintain this junction through their interaction with each other are the ONM-localized Nvj1p and the tonoplast-localized Vac8p (Kvam and Goldfarb, 2004). Targeting to and specifically interacting with Nvj1p is the soluble oxysterol binding protein Osh1. The Arabidopsis homolog for this protein is At4g12460 (Jouhet et al., 2006). While the functions of this protein still need to be investigated in plants, the yeast homologue of Osh1 is known to bind oxysterol, an oxygenated cholesterol derivative, is linked to post-synthetic regulation of sterols and mediates lipid exchange at membrane contact sites like the nucleus-vacuole junctions (Kvam and Goldfarb, 2004; Jouhet et al., 2006).
The nucleus-vacuole junction is implied in regulating lipid homeostasis of the NE by a process called piecemeal microautophagy of the nucleus (PMN). During PMN, small portions of the nucleus, including membranes and nuclear contents, are pinched off into the vacuolar lumen by the nucleus-vacuole junctions for degradation. Localization of Osh1 at these junctions is important for this process suggesting that oxysterol is here used as a signalling messenger to induce PMN (Kvam and Goldfarb, 2004). This form of autophagy has been suggests to be a ‘house-cleaning’ mechanism for the nucleus under both normal and stress conditions by clearing out excess protein and membrane as well as domains of the nucleaus that are damaged or no longer active (Mijalijcs et al., 2010). Again, this process has been well studied in yeast and is also known to occur in mammalian systems. Whether it occurs in plants is so far not known. However, microautophagy certainly takes place in plant cells and the need to regulate the composition of the NE and nucleus per se as well as the presence of a plant Osh1 homologue indicate that a process like PMN could also take place in plants.
2.10.2 NE Phospholipid Regulation by Lipins
Apart from trafficking phospholipids from their place of synthesis, the ER, to the NE, to maintain the composition of the NE membranes, metabolism of phospholipids can also occur directly at the NE. Enzymes responsible for this are phosphatases called lipins and they are involved in two different processes (Sinissoglou, 2009). Firstly, they dephosphorylate phosphatidic acid into diacylglycerol and are thus involved in the triacylglycerol synthesis and phospholipid metabolism. Secondly, they are implied in directly regulating expression of genes involved in lipid metabolism. In addition, the yeast lipin Pah1 is also required for maintaining the spherical shape of nuclei (Sinissoglou, 2009). Knock-outs of Pah1 result in irregularly shaped nuclei with long stacks of NPC-containing membranes associated with the NE. As this soluble protein is translocated into the ONM, it is thought that changes in the ONM phosphatidic acid homeostasis in the knock out lead to the NE expansion (Sinissoglou, 2009). Two lipin homologues have been identified in Arabidopsis and named AtPah1 and AtPah2 (Nakamura et al., 2009). Like their yeast and mammalian counterparts, they contain phosphatase activity and are essential for maintaining lipid metabolism and adaptation to phosphate starvation. During phosphate starvation in plants, membrane lipid remodelling occurs, whereby phospholipids are converted to non-phosphorus galactolipids to free up phosphorus. In AtPah1/AtPah2 double knock-out plants this remodelling is severely compromised (Nakamura et al., 2009). However, so far it remains unclear whether either of the two proteins is localized in the plant NE and whether a single or double knock-out affects NE structure and function. Irrespective of this, it is becoming clear that coordinating production, transport and remodelling of phospholipids is essential in maintaining the structure and function of eukaryotic membranes including the plant NE.
2.11 The Role of Plant NE Components in Stress Responses
2.11.1 Nuclei Repositioning in Response to Environmental Stimuli
It was already mentioned (Section 2.5) that cytoskeletal associations with the NE are involved in nuclear movement due to abiotic and biotic stress. Indeed, nuclear repositioning is one of the earliest defensive cell responses to external stimulation. Abiotic stimuli studied so far include blue light exposure and mechanical stress. In Arabidopsis leaf cells, nuclei positioning depends on light exposure. In the dark, nuclei are located at the bottom centre of the cell adjacent to the periclinal wall. Upon exposure of strong blue light of 50 μmol m−2s−1 and more the nuclei move to the anticlinal wall. This repositioning event is thought to protect from UV-induced DNA damage. Instead of being caused by general cytoplasmic streaming, the light induced nuclei repositioning is specifically regulated by phototropin receptor 2 and involves significant reorganization of the actin cytoskeleton (Iwabuchi et al., 2007; Iwabuchi et al., 2010).
In addition to light sensitivity, nuclear movement has also been observed in response to mechanical stimulation. Plant cells are exposed to repeated and varied strength mechanical stimulation exerted from neighbouring cells as well as from pathogens such as fungal hyphae. It has been shown that nuclei are highly sensitive to such pressures and can react to repeated stimuli of various strength by moving towards the sites of the stimuli without any loss of velocity or sensitivity (Figure 2.6; Qu and Sun, 2007). Pressures exerted from neighbouring cells are due to growth and cell division. In fact, the intracellular position of the nucleus predefines the division plane, hence appropriate nuclear positioning is essential for correct proliferation and morphology of plant tissues and organs (Qu and Sun, 2008).
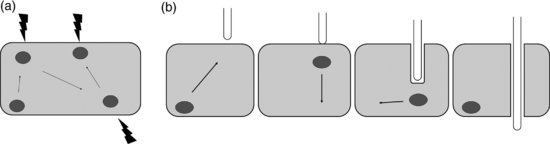
While is has been shown that elements of the cell wall, plasma membrane and microtubule cytoskeleton are responsible for transduction of mechanical signals (Qu and Sun, 2008), it is reasonable to speculate that components of the NE are also involved in these signalling events, which may effect NE-cytoskeletal associations, which in turn facilitate the nuclear movement. It will be a significant contribution to study the role of NE components in orchestrating nuclear repositioning events.
More is know about the function of NE components in response to fungal and bacterial pathogens. Plants undergo symbiotic relationships with both fungi and bacteria to effectively take up phosphorous and nitrogen, respectively. Medicago truncatula is a model organism to study both of these symbiotic relationships. Gigaspora is an arbuscular mycorrhizal fungus, which infects M. truncatula roots. During this infection, fungal hyphae penetrate root epidermal cells to spread towards the inner root cortex. The cell penetration is controlled by both plant and fungus and the plant nucleus plays a crucial role by orchestrating cellular events of this process (Genre et al., 2005). As the hyphae exert pressure onto the epidermal cell, the nucleus moves towards this area (Figure 2.6), called an appressorium. Cytoskeletal elements and ER membranes accumulate in the space between nucleus and appressorium and together with the nucleus form the pre-penetration apparatus (Genre et al., 2005). This assembly step is followed by the nucleus moving away from the appressorium towards the opposite side of the wall at speeds of 15–20 μm/h (Figure 2.6). Following the moving nucleus, endomembranes, actin and microtubule cytoskeleton reorganize to initiate the formation of a cytoplasmic column that stretches between the appressorium and the nucleus. It is inside this cytoplasmic column that the penetrating fungal hyphae traverses the plant cell (Genre et al., 2005). While NE components associated with the cytoskeleton may be hypothesized to function in the nuclear movement, it has been shown that the NE-localized ion channel DMI1 is involved in Ca2+ signalling that governs this process (Genre et al., 2005; Chabaud et al., 2011).
This DMI1-dependent Ca2+ signalling has also been shown to be required for nodulation upon rhizobial bacteria infection of M. truncatula (Riely et al., 2006; Peiter et al., 2007). The bacteria release Nod factors, which trigger calcium spikes associated with the nucleus of the infected root cell. These calcium signals, in turn, activate the expression of genes involved in formation of root nodules. The NE- localized DMI1 (Riely et al., 2006) is required for the generation and control of the calcium spikes but whether it acts as a Ca2+ or other cation channel remains unclear (Peiter et al., 2007; Matzke et al., 2009). Rhizobial bacteria enter the root system through infection threads that stretch from the root hair to other root cell layers. Similar to the cytoplasmic tubes that contain the penetrating fungal hyphae these infection threads require the reorganization of cytoskeletal elements and endomembranes. Notably, the growing infection thread in the root hair follows at a set distance to the migrating nucleus, which moves towards the bottom of the cell (Gage, 2004) suggesting that here, too, the nucleus marks the path of infection threads.
2.11.2 Functions of the Plant NE during Viral Infection
Apart from fungal and bacterial infection, the nucleus also moves towards the site of viral infection. This has been studied in Vigna unguiculata (cowpea) during infection with cowpea rust virus. Similar to other nuclear movement events, this repositioning is actin dependent (Skalamera and Heath, 1998). During viral infection, NE components may not only play a role in nuclear movement, but the NE itself is utilized by the virus for replication and formation of mature virions. Such events have been studied during barley yellow dwarf virus (BYDV) and Sonchus yellow net virus (SYNV) infections (Dennison et al., 2007; Goodin et al., 2007). Both are RNA viruses whose replication cycles include a nuclear stage. The BYDV has a movement protein that is required for the transport of viral RNA (vRNA) into the nucleus. The exact mechanism remains to be studied, but it was found that this movement protein associates with and causes conformational changes of the NE. The N-terminus of this protein can directly interact with lipids and causes the formation of protrusions that emanate from the NE (Liu et al., 2005). The lipid interaction is probably mediated by an amphiphilic α-helix at the N-terminus (Liu et al., 2005; Dennison et al., 2007). In addition, it was found that the movement protein can alter the physical properties of the NE, specifically its fluidity, and thereby induce surface pressures of as much as 2 mNm−1 (Dennison et al., 2007). These alterations of the NE are thought to be involved in integrating the protein into the membrane. The purpose of this remains hypothetical with suggestions that it is either involved in the translocation of vRNA across the NE membranes (Liu et al., 2005) or targets and associates vRNA at the NE (Dennison et al., 2007).
By contrast, during SYNV infection, the ribonucleoprotein core of the virus is trafficked into the nucleus via the NPC (Jackson et al., 2005). SYNV infection also causes NE morphological changes – nuclei are larger and numerous intranuclear membrane protrusions have been observed. Goodin et al. (2007) found that these protrusions are extensions of the NE and involved in maturation of virions. The SYN virus consists of a ribonucleoprotein core that is surrounded by a membrane layer. The membrane contains a viral glycoprotein and is thought to be attached to the viral core by a viral matrix protein. Viral replication and assembly of the core occurs inside the nucleus. Using fluorescent fusions to the matrix and glycoprotein, Goodin et al. (2007) have shown that the glycoprotein is located in the NE and the matrix protein associated with the nuclear side of the NE. The assembled core virions are thought to bud through the INM at the intranuclear membrane extensions. By budding through the INM, the virions acquire their membrane coat. Once in the periplasm, it is thought that the mature virions move into the ER lumen and traffic throughout the ER to either the cell periphery or plasmodesmata for cell-to-cell infection (Goodin et al., 2007). The formation of the intranuclear membranes and the presence of viral glycoprotein in the plant NE demonstrate that the viral infection interferes with nucleoskeletal and NE membrane components to effect such dramatic changes.
2.12 Concluding Remarks
The NE is an important structure as it is essential for a variety of cellular and nuclear processes. We have only just caught a glimpse of the functional capacity of the plant NE with many questions remaining to be answered. What are the protein components of the plant NE and what are their functional roles? Why is the plant NE structurally similar to yeast and metazoan NEs, yet seems to contain plant-specific components? How is the NE involved in chromatin anchorage and organization? What molecular processes define the roles the NE plays in stress and disease responses? The field of plant NE biology should have a bright future with a lot of potential for novel and exciting research.
Acknowledgements
DE and KG Acknowledge the support of the Leverhulme Trust under grant F/00 382/H and through a Leverhulme Early Career Fellowship to KG.